1Department of Microbiology, Faculty of Science, Annamalai University,
Annamalai Nagar, Chidambaram, Tamil Nadu, India.
2Department of Plant Science, Avvayar Government College for Women,
Karaikal, Puducherry, India.
Corresponding author email: sivasubramanik@gmail.com
Article Publishing History
Received: 10/07/2021
Accepted After Revision: 15/09/2021
In our going research on identifying interesting bioactive molecules from various natural resources, marine microbes represent an interesting category because of their large biodiversity and unexploited background to the scientific world. Many of the marine natural products for those particularly procured from macroorganisms, have already undergone clinical trials. So, the research on natural products derived from marine microbes have tremendously increased in the recent years due to the demand of bioactive molecules having pharmaceutical potential for various target applications. Hence, the present study was undertaken with an aim to purify and chemically characterize a potential bioactive molecule from a symbiotic marine bacterium, Paenibacillus macerans SAM 9 isolated from a sea anemone, Heteractis aurora. After microbial fermentation under standardized conditions, the bioactive molecule was ethyl acetate extracted and purified using a Reverse Phase-HPLC method.
The purified fraction revealed promising antimicrobial activity against a clinical human pathogen, Proteus mirabilis which was extensively studied for the chemical and structural characterization. Using FT-IR, 1H and 13C NMR, GC and MS/MS spectral studies, the bioactive molecule was identified as (E)-10-hydroxy-4,6,8,10-tetramethyldodec-4-en-3-one. This is the first report that the (E)-10-hydroxy-4,6,8,10-tetramethyldodec-4-en-3-one is identified from a bacterium and from this study, this bioactive molecule can easily be procured using the standardized conditions explored in this investigation. Moreover, this study acts as the baseline data for the purification and future exploration of this bioactive molecule regarding its extensive biomedical significance. The aim of this study is to become a suitable milestone and act as a credible source for future studies.
Bioactive, (E)-10-Hydroxy-4,6,8,10-Tetramethyldodec-4-En-3-One,
Marine Symbiotic Bacterium, RP-HPLC, Spectral Studies
Bharathi T, Sambandan K, Sivasubramani K. Structural Characterization of (E)-10-Hydroxy-4,6,8,10-Tetramethyldodec-4-en-3-one Bioactive Molecule from Marine Paenibacillus macerans Isolate. Biosc.Biotech.Res.Comm. 2021;14(3).
Bharathi T, Sambandan K, Sivasubramani K. Structural Characterization of (E)-10-Hydroxy-4,6,8,10-Tetramethyldodec-4-en-3-one Bioactive Molecule from Marine Paenibacillus macerans Isolate. Biosc.Biotech.Res.Comm. 2021;14(3). Available from: <a href=”https://bit.ly/3k8s6s9“>https://bit.ly/3k8s6s9</a>
Copyright © Bharathi et al., This is an open access article distributed under the terms of the Creative Commons Attribution License (CC-BY) https://creativecommns.org/licenses/by/4.0/, which permits unrestricted use distribution and reproduction in any medium, provide the original author and source are credited.
INTRODUCTION
Presently many bioactive compounds have been derived from various terrestrial resources. Hence, the exploration of interesting, novel and needful biochemical metabolites to the biomedical applications is becoming low (Jensen and Fenical 2000). So, the need of new sources for the exploration of bioactive substances is much required nowadays. In this condition, marine ecosystem offer enormous biodiversity and have potential resources to find interesting bioactive molecules with numerous worthwhile applications.
As per the Global Biodiversity Assessment by the United Nations Environment Programme, there are 178,000 marine species belonging to 34 phyla have been documented which reveals that the ocean’s biodiversity represents 50% of the globe’s whole biodiversity and making marine microbes a sustainable and appreciable source of interesting bioactive molecules discovery (Jimeno et al. 2004; Balan et al. 2012; Mitra and Zaman 2016; Balan et al. 2019b; Yadav 2021). Many of the marine natural products for those particularly procured from macroorganisms, have already undergone clinical trials (Newman and Cragg 2004; Yadav 2021).
So, research on natural products derived from marine microbes have tremendously increased in the recent years due to the demand of bioactive molecules having pharmaceutical potential for various target applications (Andersen and Williams 2000; Balan 2014). In contrast to macroorganisms, microorganisms represent appreciable natural product resources having the advantage of feasible and sustainable production of secondary metabolites under large quantities at reasonable cost as well as the fermentation of the whole microbial species (Waites et al. 2009; Balan et al. 2013).
Further, adaptive species of the marine ecosystem characterize very special and unique conditions which differ them from those found in other habitats. Furthermore, marine microbes sometimes accumulate structurally unique bioactive molecules which are not relatively present in terrestrial sources (Bhakuni and Rawat 2006; Balan and Jayalakshmi 2013; Balan et al. 2019a; Varijakzhan et al. 2021).
Considering the above facts, the present study aims to identify the bioactive molecule produced by a marine symbiotic bacterium, Paenibacillus macerans SAM 9. Further, this strain has already been reported for its potential antimicrobial activity against a panel of clinical human pathogens in our previous study and this present study structurally characterize the molecule responsible for the bioactivity through a systematic scheme of spectral characterization after purification (Bharathi et al. 2021). To the best of our knowledge, Paenibacillus macerans have never been reported earlier for the identification and its detailed structural characterization of a bioactive molecule. Hence, exploring the chemical nature of this bioactive molecule makes sense in regards with the novelty and this warrants the present investigation.
MATERIAL AND METHODS
For the symbiotic marine bacterium and culture conditions, Paenibacillus macerans SAM 9 is the bioactive molecule producing symbiotic marine bacterium used in this study which was previously isolated from the sea anemone, Heteractis aurora and the 16S rRNA partial sequence of this bacterium was submitted in the NCBI GenBank with the accession no. MT941031.1 (Bharathi 2021). In our earlier study, this marine bacterium was reported to produce a bioactive molecule which exhibited promising antimicrobial activities against many clinically isolated human pathogens.
The fermentation production of this bioactive molecule was carried out here for the purification and detailed structural characterization using a previously optimized microbiological medium intended for the maximum production of this bioactive compound with the following conditions: 1% maltose, 0.5% ammonium nitrate, 34 ppt salinity, pH 8, 35°C temperature with an incubation period of 96 hrs, respectively. Further, the inoculum was prepared using 1% exponential phase culture in the same fermentation medium as described above, where the optical density (OD620 nm) of the inoculum was adjusted to 0.1 based on McFarland turbidity standard of 0.5, equivalent to the bacterial concentration of 1×108 cfu ml−1. The fermentation was carried out in one litre conical flask with 400 ml working volume.
The extraction and HPLC purification of the bioactive molecule was done using optimized conditions. Production of bioactive molecule was carried out freshly and the bioactive molecule was extracted from the cell free supernatant using ethyl acetate after centrifugation at 3000 rpm for 30 min. After an overnight incubation, the organic phase containing the bioactive molecules was separated and rotary vacuum evaporated at 50ºC to a dried form. The dried form was subjected to purification of this bioactive molecule using a high-pressure liquid chromatography (HPLC). The dried crude form was initially dissolved in 3:2 ratio of 5 ml acetonitrile and methanol and passed through a 0.2 μm syringe filter.
The purification was done by Reverse Phase (RP)—C18 silica gel (230–400 mesh) column at 30 °C temperature. The solvent system consisted of acetonitrile (solvent A) and methanol (solvent B) and the elutions were made at 0.5 ml/min flow rate using the stepwise gradient initiated from the ratio of 60:40, vol/vol (A:B) to the finalized ratio of 100:0, vol/vol (A:B). The absorbance of the eluate was measured at 210 nm. All the collected fractions were dried individually using rotatory vacuum evaporation and studied for the antimicrobial activity.
For antimicrobial assay, all the dried fractions of the HPLC purification were individually studied for antimicrobial activity using the most susceptible pathogen, Proteus mirabilis of this bioactive molecule as reported in our earlier study. The human clinical pathogen, P. mirabilis was kindly gifted by Rajah Muthiah Medical College Hospital, Annamalai University, Tamil Nadu, India. The antimicrobial activity of the purified fractions was evaluated using microtiter plate-based assay method (Casey et al. 2004).
The assay was conducted in a 24-well flat bottom polystyrene microtitre plates with lids (Tarsons, India). Briefly, the well plates were filled with 100 μL of sterile tryptone soy broth followed by the addition of 10 μL purified (dried) fraction dissolved in phosphate buffer saline (PBS: 10 mM KH2PO4/K2HPO4 and 150mM NaCl with pH adjusted to 7.0) and 10 μL of the pathogenic P. mirabilis culture. The growth control well plates was done by the addition of 100 μL of tryptone soy broth, 10 μL pathogenic P. mirabilis culture and 10 μL phosphate buffer saline (without any addition of purified fraction).
The susceptibility control well plate was carried out with 100 μL of tryptone soy broth, 10 μL pathogenic P. mirabilis culture and 10 μL of 4mg/ml streptomycin (commercial antibiotic). After inoculation, the well plates were covered with lids and incubated at 37 °C for 48 hrs. After incubation, the absorbance of the well plates was recorded at 600 nm using a microplate reader (Biotek Elx808, WI, USA), where the assays were carried out in triplicate. The growth inhibition percentages of the individual purified fractions were calculated as follows:
% Growth inhibition = [(1 − (As/Ac)] × 100
Where As represents the absorbance of the well with purified fractions and Ac represents the absorbance of the growth control well (without any added bioactive sample). The fraction showed the appreciable antimicrobial activity was studied for further detailed chemical and structural characterization of the bioactive molecule. For the chemical and structural characterization of purified bioactive molecule, the purified fraction showing antimicrobial activity against the clinical human pathogen P. mirabilis was identified for the bioactive molecule present in it. The compound was chemical analyzed and structurally characterized as per the following instrumentation procedures.
Fourier Transform Infrared Spectroscopy (FT-IR) was used to identify the characteristic functional groups of the bioactive molecule present in the purified bioactive fraction (Mani et al. 2016a). Five milligram of the purified molecule was dispersed in dry powdery potassium bromide (KBr), the resultant was thoroughly mixed using a mortar and pressed at 6 bars pressure within 2 min to form a KBr thin disc. The resulting disc was placed in a sample cup of a diffuse reflectance accessory. Infrared absorption spectrum was analyzed on an IR affinity, FT-IR system (Shimadzu, Japan) at a spectral resolution of 4 cm-1 with an average of 10scans in the wavenumber range between 400–4000 cm-1.
1H & 13C Nuclear Magnetic Resonance Spectroscopy (NMR) spectrum of the purified bioactive molecule was analyzed using a Bruker AV600NMR spectrometer (Germany) in which the deuterated CDCl3 was used as the solvent. Chemical shifts were expressed in parts per million (ppm) downfield from an internal standard of tetramethylsilane (TMS). The purified fraction containing the bioactive molecule was characterized using a Gas Chromatography and Mass Spectroscopy (GC-MS).
GC-MS was performed on a Thermo Trace GC Ultra coupled with Polaris Q MS and TriPlus auto-sampler using a DB-5 (0.25 mm × 30 m × 0.22 μm) column in which helium was used as carrier gas. The temperature was set between 50°C to 250°C at a rate of 10°C min-1. The initial temperature was held for 2 min and final temperature of 250°C was held for 10 min. The GC flow rate was 1 ml min–1 and the total run time was 32 min. MS was performed at scan mode between 0 – 300 m/z with an Ion trap EI+.
RESULTS AND DISCUSSION
Extraction and purification of bioactive compounds are important steps as it determines the level of significance and use as well as identification of the particular molecule (Thangaraj 2016). Initially, the production of bioactive molecule by the marine symbiotic bacterium, P. macerans SAM 9 was done as per the optimized fermentation conditions from our earlier study.
Further, the ethyl acetate solvent was used for the extraction of this bioactive molecule (Fig. 1) in which organic solvent was pooled and concentrated using rotary vacuum evaporation. The concentrated dried form was further used for the purified with a HPLC and there were seven fractions obtained (Fig. 2). All the fractions were individually collected and dried with rotary vacuum evaporation and studied for the antimicrobial activity against the human clinical pathogen P. mirabilis (Thangaraj 2016).
Figure 1: Solvent extraction of the bioactive molecule produced by P. macerans SAM 9
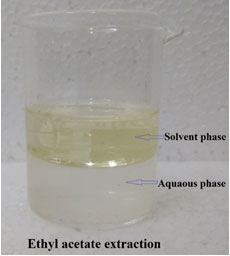

Among the fractions, fifth fraction alone showed antimicrobial activity with 85.5 ± 2.3 % which confirmed the presence of bioactive molecule present in it and rest of the fractions revealed no antimicrobial activity. The fifth fraction evidenced antimicrobial activity was subjected to further detailed chemical and structural characterization studies. Similar to this study, earlier investigations have noted that the significance of non-polar solvents followed by reverse phase column chromatography for the purification of bioactive compounds (Mani et al. 2016b; Balan et al. 2017; Balan et al. 2019b).
Figure 2: HPLC chromatogram of the crude ethyl acetate extract
from the cell free supernatant of P. macerans SAM 9
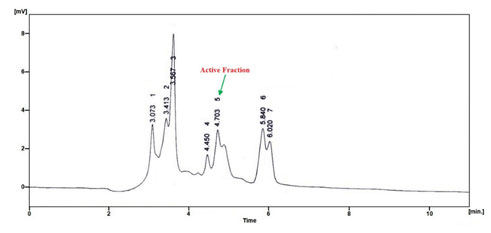
Figure 3: FT-IR Spectrum of the purified bioactive molecule,
(E)-10-hydroxy-4,6,8,10-tetramethyldodec-4-en-3-one
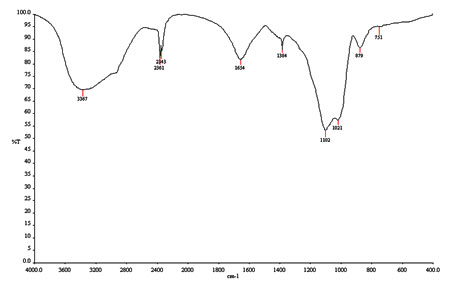
FT-IR spectrum of the purified bioactive molecule revealed all the functional groups of (E)-10-hydroxy-4,6,8,10-tetramethyldodec-4-en-3-one as shown in the Figure 3. The terminal alkane group (CH3) present at the wavelengths of 1384cm-1, similarly, intermittent alkane groups (CH2 and CH) of the bioactive molecule were observed at 751 and 1021cm-1 wavelengths. Further, the alkene group (C=CH) was depicted at wavelength of 1654 and 879cm-1. Likewise, the most significant groups of ether (C=O-CH2) were predicted at 2343 and 2361cm-1 and the aldehyde (C-OH) was evidenced at 1102 and 3367cm-1 wavelengths, respectively.
1H and 13C NMR spectrum of the purified bioactive molecule represented all the functional groups of (E)-10-hydroxy-4,6,8,10-tetramethyldodec-4-en-3-one as shown in the figures 4 and 5. 1H NMR spectrum revealed the existence of terminal alkane hydrogen (CH3) within chemical shifts of 0.8821 – 0.9674ppm, similarly, the chemical shifts between 1.0296 – 1.3532ppm and 1.4147 – 1.5908ppm responsible for the presence of aliphatic alkane hydrogen atoms (CH2 and CH). The presence of protons (hydrogen atom) in the alkene group (C=CH) evidenced from the chemical shifts between 5.5038 to 5.7548ppm. Further, the protons within the chemical shifts, 4.0532 – 4.1310ppm and 9.8021 – 9.9802ppm corresponds to the existence of aldehyde hydrogen (C-OH) and ether hydrogen (C=O-CH2) which were the significant functional groups of (E)-10-hydroxy-4,6,8,10-tetramethyldodec-4-en-3-one.
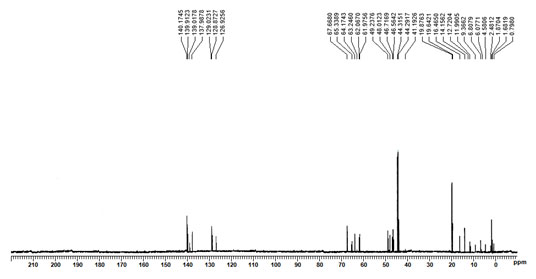
Furthermore, 13C NMR spectrum of this bioactive molecule revealed the presence of same functional groups of (E)-10-hydroxy-4,6,8,10-tetramethyldodec-4-en-3-one as predicted in the 1H NMR spectrum. The presence of terminal alkane carbon (CH3) and alkane carbons (CH2 and CH) were observed between the chemical shifts of 0.7980 – 4.5806ppm, 6.0771 – 9.3662ppm and 11.9905 – 19.8763ppm, similarly, the presence of alkene carbon (C=CH) was predicted within 126.9256 – 140.1745ppm. Moreover, the carbon atom from the significant functional groups of aldehydes (C-OH) and carbonyl (C=O) groups were represented within the chemical shifts of 61.9756 – 67.6680ppm and 41.1926 – 49.2376ppm.
Figure 4: 1H NMR spectrum of the purified bioactive compound,
(E)-10-hydroxy-4,6,8,10-tetramethyldodec-4-en-3-one
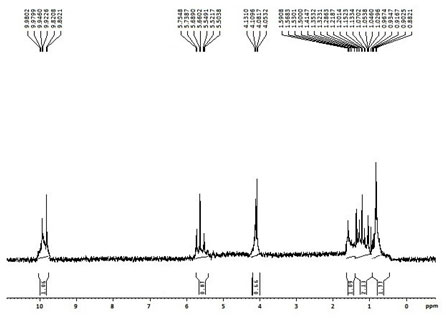
Figure 5: 13C NMR Spectrum of the purified bioactive compound,
(E)-10-hydroxy-4,6,8,10-tetramethyldodec-4-en-3-one
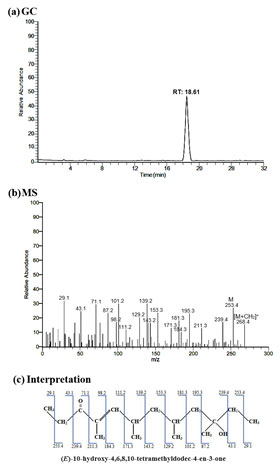
GC-MS analysis of purified bioactive molecule was illustrated in the Figure 6. GC spectrum exhibited only one molecular ion peak at the retention time of 18.61 min which revealed the presence of single bioactive molecule in the purified fraction (Fig. 6a). Further, the mass spectral analysis evidenced that the peak was (E)-10-hydroxy-4,6,8,10-tetramethyldodec-4-en-3-one with a molecular mass of methylated and non-methylated form with 268.4 and 253.3 MW (Fig. 6b).
Further, the interpretation of the mass spectrum showed exact sequential molecular weight pattern of (E)-10-hydroxy-4,6,8,10-tetramethyldodec-4-en-3-one under methylated form which were 29.1, 43.1, 71.1, 87.2, 98.2, 101.2, 111.2, 129.2, 139.2, 143.2, 153.3, 171.3, 181.3, 184.3, 195.3, 211.3, 239.4, 253.4 and 268.4 MW, respectively (Fig. 6c).
Furthermore, this molecular pattern revealed exact similarity of (E)-10-hydroxy-4,6,8,10-tetramethyldodec-4-en-3-one with the available NIST database used for the molecular identification of chemical compounds. All these above information using FT-IR, NMR and GC-MS analysis confirmed that the purified bioactive molecule was a (E)-10-hydroxy-4,6,8,10-tetramethyldodec-4-en-3-one and exact molecular structure was illustrated in the figure 7 based on the above spectral information. Similar to the present study, various earlier studies have used FT-IR, NMR and MS spectral analysis for the structural identification of different bioactive molecules.
Recently, a bioactive molecule purified from a marine yeast, Cyberlindnera saturnus SBPN named as Cybersan revealed a molecular structure of three galactose followed by a heptadecanoic acid which was active against several human clinical pathogen bacteria (Balan et al. 2019a). Likewise, Staphylosan, a bioactive glycolipid compound purified from a marine bacterium, Staphylococcus saprophyticus SBPS-15 showed a molecular structure of mannose-mannose-oleic acid which exhibited antimicrobial and biofilm activity against many marine biofilm forming bacteria and Pontifactin a bioactive lipopeptide compound identified from a marine Pontibacter korlensis SBK-47 evidenced the chemical structure of palmitic acid-serine-aspartic acid-valine-serine-serine which was biological active against many MTCC pathogenic bacterial strains (Balan et al. 2016; Balan et al. 2019b).
Figure 6: (a) GC, (b) MS and (c) Interpretation based on molecular weight pattern
of the methylated (E)-10-hydroxy-4,6,8,10-tetramethyldodec-4-en-3-one
The bioactive molecule identified in this study, (E)-10-hydroxy-4,6,8,10-tetramethyldodec-4-en-3-one was earlier isolated from an actinomycete, Actinopolyspora erythraea YIM 90600 which was named as Actinopolysporins C. Till date, this metabolite is not reported from any bacterial species, henceforth, this is the first study reporting the production of this biomolecule from a bacterium originated from marine environment (Zhao et al. 2011; Balan et al. 2019b).
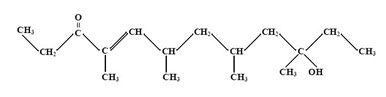
Figure 7: Structure of (E)-10-hydroxy-4,6,8,10-tetramethyldodec-4-en-3-one
CONCLUSION
The findings of the present study reveals that the bioactive molecule from the marine symbiotic bacterium, P. macerans SAM 9 was extracted with ethyl acetate and purified using a Reverse Phase C18 silica gel column. The purified bioactive molecules were identified as (E)-10-hydroxy-4,6,8,10-tetramethyldodec-4-en-3-one based on various spectral studies. The present investigation proved that this bacterium is the promising source for the appreciable production of this excellent bioactive compound, further, this study holds as the baseline data for the exploration of many biomedical properties of this bioactive molecule in near future.
ACKNOWLEDGEMENTS
This study was supported by the Department of Microbiology, Faculty of Science, Annamalai University, Chidambaram, Tamil Nadu, India. Authors thank the department for providing lab facilities.
Conflict of Interest: Authors declare no conflict of interests to disclose.
REFERENCES
Andersen, R.J. and Williams, D.E., (2000). Chemistry in the marine environment. Pharmaceuticals from the Sea. The Royal Society of Chemistry, Cambridge, UK, pp.55-79.
Balan, S.S. and Jayalakshmi, S., (2013). Glycolipid biosurfactant production using low-cost medium from marine bacterium Pseudomonas aeruginosa of Mudasalodai coast. International Journal of Green Chemistry and Bioprocess, 3(3), pp.33-37.
Balan, S.S., Kumar, C.G. and Jayalakshmi, S., (2016). Pontifactin, a new lipopeptide biosurfactant produced by a marine Pontibacter korlensis strain SBK-47: Purification, characterization and its biological evaluation. Process Biochemistry, 51(12), pp.2198-2207.
Balan, S.S., Kumar, C.G. and Jayalakshmi, S., (2017). Aneurinifactin, a new lipopeptide biosurfactant produced by a marine Aneurinibacillus aneurinilyticus SBP-11 isolated from Gulf of Mannar: Purification, characterization and its biological evaluation. Microbiological Research, 194, pp.1-9.
Balan, S.S., Kumar, C.G. and Jayalakshmi, S., (2019a). Physicochemical, structural and biological evaluation of Cybersan (trigalactomargarate), a new glycolipid biosurfactant produced by a marine yeast, Cyberlindnera saturnus strain SBPN-27. Process Biochemistry, 80, pp.171-180.
Balan, S.S., Mani, P., Kumar, C.G. et al. (2019b). Structural characterization and biological evaluation of Staphylosan (dimannooleate), a new glycolipid surfactant produced by a marine Staphylococcus saprophyticus SBPS-15. Enzyme and Microbial Technology, 120, pp.1-7.
Balan, S.S., Nethaji, R., Sankar, S. et al. (2012). Production of gelatinase enzyme from Bacillus spp isolated from the sediment sample of Porto Novo Coastal sites. Asian Pacific Journal of Tropical Biomedicine, 2(3), pp.S1811-S1816.
Balan, S.S., Raffi, S.M. and Jayalakshmi, S., (2013). Probing of potential luminous bacteria in Bay of Bengal and its enzyme characterization. Journal of Microbiology and Biotechnology, 23(6), pp.811-817.
Balan, S.S., (2014). Production, characterization, evaluation of a glycolipid biosurfactant from a marine strain Bacillus cereus and development of non-toxic skin and hair care cosmetic formulations, Doctoral dissertation, Annamalai University, Tamil Nadu, India, 63-98.
Bhakuni, D.S. and Rawat, D.S., (2006). Bioactive marine natural products. Springer Science & Business Media.
Bharathi, T., Sambandan, K. and Sivasubramani, K., (2021). Enhanced production of a bioactive molecule from a symbiotic marine bacterium, Paenibacillus macerans SAM 9 isolated from the sea anemone, Heteractis aurora. Journal of Applied Biology and Biotechnology, 9(1), pp.17-24.
Casey, J.T., O’Cleirigh, C., Walsh, P.K. et al. (2004). Development of a robust microtiter plate-based assay method for assessment of bioactivity. Journal of Microbiological Methods, 58(3), pp.327-334.
Jensen, P.R. and Fenical, W., (2000). Marine microorganisms and drug discovery: current status and future potential. Drugs from the Sea, pp.6-29.
Jimeno, J., Faircloth, G., Sousa-Faro, J.M., et al., (2004). New marine derived anticancer therapeutics─ a journey from the sea to clinical trials. Marine Drugs, 2(1), pp.14-29.
Mani, P., Dineshkumar, G., Jayaseelan, T., et al., (2016a). Antimicrobial activities of a promising glycolipid biosurfactant from a novel marine Staphylococcus saprophyticus SBPS 15. 3 Biotech, 6(2), pp.1-9.
Mani, P., Sivakumar, P. and Balan, S.S., (2016b). Economic production and oil recovery efficiency of a lipopeptide biosurfactant from a novel marine bacterium Bacillus simplex. Achievements in the Life Sciences, 10(1), pp.102-110.
Mitra, A. and Zaman, S., (2016). Basics of marine and estuarine ecology. Springer, pp.53-87. https://doi.org/10.1007/978-81-322-2707-6.
Newman, D.J. and Cragg, G.M., (2004). Marine natural products and related compounds in clinical and advanced preclinical trials. Journal of Natural Products, 67(8), pp.1216-1238.
Thangaraj, P., (2016). Extraction of bioactive compounds. In Pharmacological assays of plant-based natural products, Springer, Cham, pp. 11-13.
Varijakzhan, D., Loh, J.Y., Yap, W.S., et al., (2021). Bioactive Compounds from Marine Sponges: Fundamentals and Applications. Marine Drugs, 19(5), p.246. https://doi.org/10.3390/md19050246.
Waites, M.J., Morgan, N.L., Rockey, J.S., et al., (2009). Industrial Microbiology: An introduction. John Wiley & Sons.
Yadav, A.N., 2021. Microbial biotechnology for bio-prospecting of microbial bioactive compounds and secondary metabolites. Journal of Applied Biology & Biotechnology, 9(2), pp.1-6.
Zhao, L.X., Huang, S.X., Tang, S.K., et al., (2011). Actinopolysporins A–C and tubercidin as a Pdcd4 stabilizer from the halophilic actinomycete Actinopolyspora erythraea YIM 90600. Journal of Natural Products, 74(9), pp.1990-1995.


