Department of Microbiology, Ch. Charan Singh University, Meerut- 250004, India
Vice-Chancellor, Shobhit Institute of Engineering & Technology (Deemed-to-be University), NH-58, Modipuram, Meerut- 250110, India”
Corresponding author email: amarprakashgarg@yahoo.com
Article Publishing History
Received: 09/10/2019
Accepted After Revision: 30/11/2019
In the present study we have evaluated the antagonistic spectrum of 8 isolates of Lactobacillus against common enteric pathogens followed by auto-aggregation, co- aggregation and cell surface hydrophobicity. Isolate C9 showed antagonistic activity against all test species namely Bacillus subtilis, Bacillus cereus, Salmonella enteric, Shigella flexneri, Streptococcus pneumonia , Staphylococcus aureus , Staphylococcus epidermidis, Pseudomonas aeruginosa, E. coli, Clostridium perfringens, Listeria monocytogens and yeast Candida albicans that were obtained from IMTECH, Chandigarh. G4 showed highest zone of inhibition against Listeria monocytogens (20mm) while C28 exhibited highest zone of inhibition against Shigella flexneri. Similarly P37 showed against Salmonella enteric (24mm); C9 showed highest zone of inhibition against Pseudomonas aeruginosa (20mm) and E. coli (22mm). All the 8 Lactobacillus isolates exhibited the remarkable inhibitory effects against all test pathogenic strains with variable spectrum of inhibition. C9 showed highest auto-aggregation ability (91.6%) and co- aggregation activity against all the pathogens. Amongst all the eight isolates C9 showed noticeable higher hydrophobicity, hence the results suggest that the isolates Lactobacillus may be used as natural bio-preservatives in different food products and also to extent the shelf life of food products.
Pathogenic bacteria, Lactobacillus, antagonistic spectrum, bio-preservatives, shelf life.
Bisht N, Garg A. P. Antagonistic Activity of Lactic Acid Bacteria Against Common Enteric Pathogens Isolated from Milk and Milk Products and Evaluation of their Probiotic Attributes. Biosc.Biotech.Res.Comm. 2019;12(4).
Bisht N, Garg A. P. Antagonistic Activity of Lactic Acid Bacteria Against Common Enteric Pathogens Isolated from Milk and Milk Products and Evaluation of their Probiotic Attributes. Biosc.Biotech.Res.Comm. 2019;12(4). Available from: https://bit.ly/2qCpoDe
Copyright © Bisht and Garg This is an open access article distributed under the terms of the Creative Commons Attribution License (CC-BY) https://creativecommns.org/licenses/by/4.0/, which permits unrestricted use distribution and reproduction in any medium, provide the original author and source are credited.
INTRODUCTION
Increasing number of food borne pathogens and their entero- toxins has been concerned as an important research topic in the field of food safety and regulatory agencies (Kermanshahi and Qamsari, 2015). Infectious diseases caused by food borne pathogens which cause food spoilage such as Escherichia coli O157: H7, Salmonella, Listeria monocytogenes and Campylobacter cause various illness and death due to diarrhea (CDC, 2013; CDC, 2014; Dickinson and Surawicz, 2014). Commonly used antibiotics are efficient in limiting the growth of food borne pathogens, but led to expansion of bacterial resistance to antibiotics have been reported by Oroojalian et al., 2010; Andersson et al., 2010 and Campana et al., 2017 therefore the demand of new type of efficient and safe antimicrobial compounds are increasing (Gaspar et al., 2018; Bah et al., 2019).
LAB’s are potential microorganisms associated with fermentation. These are non spore forming gram positive bacteria, widely distributed in nature. Lactic acid is the main product, produced as they ferment carbohydrates primarily glucose in the raw materials to produce various metabolites which give food its unique flavor and increase nutritional value of the fermented food, which are not present before (Ren et al., 2017). LAB as natural preservatives has a wide range of antimicrobial effects (Østergaard et al., 2014 and Mejlholm and Dalgaard, 2015) against many food borne pathogens (Sharma et al., 2017; Yang et al., 2017 and Zhang et al., 2018), widely used as bio-preservatives, extend shelf life which controlled food borne pathogens and has found application in many industries and also in various commercial purposes (Adeyemo et al., 2018). The mechanism underlying the activity of LAB strains against bacterial pathogens is due to the production of bacteriocin, organic acid, ethanol, hydrogen peroxide, diacetyl and reuterin , which stimulate immune system and modulate intestinal microbiota (Tesfaye, 2014 ; Kang and Im, 2015; Bah et al., 2019).
LAB can prevent the adhesion of pathogens by competing for binding site on intestinal epithelial cell, reduced the colonization of pathogens and thereby preventing the onset of infection (Wang et al., 2018; Gao et al., 2019). Acid production is one of the mechanism by which LAB inhibit pathogens, bacteria are inactivated as the pH gets lower (Guo et al., 2017 and Wemmenhove et al., 2017). Theory of weak acids are also important, lipophilic when not dissociated; thus enter a bacterial cell through the plasma membrane and decompose into ions in a high pH environment which cause acidification of the cytoplasm (Gao et al., 2019).
Acidification alter the cell metabolism by damaging enzymes, inhibit protein synthesis, destroy genetic material, interrupt nutrient absorption and damaging the structure and function of membranes and cell wall (Hu et al., 2017). Zhang et al., 2011 proposed that aggregation of acid ions in intracellular space is important in determining antibacterial action of organic acid. Bacteriocin are ribosomally synthesized, extraecllularly released bioactive complex peptides that have a bactericidal or bacterio-static effects on other species (Masuda et al., 2012 and Costa et al., 2019). Bacteriocins have similar mechanism as that of probiotics. Bacteriocins compete with pathogenic agents for adhesion sites on mucosa (Simova et al., 2008; Jandaik et al., 2013 and Costa et al., 2019).
Bacteriocins modify the surrounding environment by modulating the pH or the oxidation-reduction potential that compromise the ability of pathogens to become established (Feliatra et al., 2018). Bacteriocin provides beneficial effects by stimulating the humoral and cellular immune response (Dhanasekaran et al., 2010; Sieladie et al., 2011 and Wang et al., 2018). Most of the produced bacteriocins by LAB are cationic peptides at a neutral pH, hydrophobic in nature and amphiphilic which contain 20 to 60 amino acids (Yang et al., 2012; Sari et al., 2018 and Costa et al., 2018). The activity of bacteriocin are related to these properties when acting on the cytoplasmic membrane where the positively charged proteins bind to negatively charged phospholipids that make up a part of the membrane of sensitive cells (Cotter et al., 2005; Kumari et al., 2008 and Yang et al., 2012 ). Bacteriocins act by creating pores in the membrane of the target cells that produces harmful effects like dissipation of proton motive force, ATP depletion and leakage of nutrients and metabolites (Costa et al., 2018 and Feliatra et al., 2018). The pore size varies from one bacteriocin to another bacteriocin according to their size, stability and conductivity (Jolly et al., 2002). Bacteriocin has bactericidal or bacteriostatic properties against spoilage and pathogens bacteria and is used as a bio-preservative in food products and also considered an additional safety measure to minimally processed products (Kumari et al., 2008; Hwanhlem et al., 2015 and Astó et al., 2019).
Intake of probiotics stimulates the growth of beneficial microorganisms which simultaneously reduces the amount of pathogens, thus improving the intestinal microbial balance and lowering the risk of gastro-intestinal diseases (Fuller, 1989; Cross, 2002; Chiang and Pan, 2012; Molina et al., 2012 and Wang et al., 2018). Probiotics are reported to also have anti-mutagenic, anti-carcinogenic, hypo-cholesterolemic, anti-hypertensive, anti-osteoporosis and immune-modulatory effects (Sieladie et al., 2011; Chiang and Pan, 2012; Shimizu et al., 2015; Wang et al., 2018; Colomer et al., 2019 and Tankoano et al., 2019). Lactic acid bacteria (LAB) is Generally Recognized as Safe (GRAS) by WHO, plays an important role in the process of fermentation of food by inhibiting spoilage bacteria and production of flavour, aroma, and texture of fermented food (Akkoc et al., 2011; Colomer et al., 2019 and Tankoano et al., 2019). The main objective of this study is to check the antagonistic activity of Lactic acid bacteria against food borne pathogens.
MATERIAL AND METHODS
Isolation of bacteriocin producing isolates
LAB strains were screened from traditional milk and milk products samples collected from different regions of Meerut. The bacteriocin producing strains from traditional milk products were isolated by the direct plating method (Coventry et al., 1997; Millette et al., 2007 and Gaspar et al., 2018) with slight modification: each sample were serially diluted ten folds with saline solution (0.9% NaCl). Aliquots (1ml) were plated onto MRS agar medium then incubated at 35 ±1°C under anaerobic conditions. Single isolates from these plates were each cultured in 10 ml of lactobacilli MRS broth for 24 h at 35 ±1°C and tested for antibacterial activity using agar well diffusion against pathogens i.e Bacillus subtilis MTCC441, Bacillus cereus MTCC430, Salmonella enteric MTCC1166, Shigella flexneri MTCC1457, Streptococcus pneumonia MTCC4673, Staphylococcus aureus MTCC7443, Staphylococcus epidermidis MTCC435, Pseudomonas aeruginosa MTCC4673, E. coli MTCC40, Clostridium perfringens MTCC 450, Listeria monocytogens MTCC 657 and yeast Candida albicans MTCC1637.
Identification of LAB
Gram’s staining, endospore staining, catalase and oxide test, arginine hydrolysis and sugar fermentation (Arabinose, Cellobiose, Galactose, Glucose, Lactose, Maltose, Mannitol, Raffinose, Ribose, Sorbitol, Sucrose and Xylose) were conducted as a preliminary steps in characterization of Lactobacillus. In order to precisely identify the species various enzymatic activities- amylases, lipases, phytases, proteases and gelatinases were done followed by probiotics attributes (acidic, bile and pancreatin) and haemolysis.
Preparation of Cell free Supernatants (CFS)
One liter of Lactobacilli MRS broth was inoculated with 10 ml of eight Lactobacillus species and incubated for 24 h at 35±1°C in bacteriological incubator (REMI). CFS was obtained by centrifugation at 10,000g for 10 min followed by neutralization of pH to 6.5 with the addition of 5 mol /l of NaOH. The resulting CFS was then filtered through a membrane filter (0.22 μm pore size).
Antimicrobial spectrum
The antimicrobial spectra of eight LAB against pathogenic bacteria such as Bacillus subtilis MTCC441, Bacillus cereus MTCC430, Salmonella enteric MTCC1166, Shigella flexneri MTCC1457, Streptococcus pneumonia MTCC4673, Staphylococcus aureus MTCC7443, Staphylococcus epidermidis MTCC435, Pseudomonas aeruginosa MTCC4673, E. coli MTCC40, Clostridium perfringens MTCC 450, Listeria monocytogens MTCC 657 and yeast Candida albicans MTCC1637 were examined by agar well diffusion assay ( Schillinger and Lucke, 1989; Batdroj et al., 2006 and Gaspar et al., 2018). From Nutrient broth 100 µl of 24 h old cultures of the pathogenic bacteria were swab on the Müller-Hinton agar (MHA) plates and afterward wells were made using sterile cork borer. Wells were filled with 100 µl of the supernatant of each isolate and plates were incubated at 35±1°C for 24 h. The diameter of inhibition zones were measured and mean diameter for the inhibition zone was recorded, standard deviation was also calculated. The test was performed in triplicate (Putra et al., 2017).
Auto-aggregation assay
For auto-aggregation assay (Ramos et al., 2013), LAB’s were grown in MRS broth for 18h at 35±1°C the cells were harvested at 9,000×g for 10 min at room temperature by centrifugation. The pellet washed twice in phosphate buffered saline (PBS) and re-suspended in PBS solution to a final concentration of about 108 cfu/ml. at this point, an absorbance was measured at 600nm (A0h) and then, 2 mL bacterial suspension were vortexed for 10s and incubated at 35±1°C for 5 h. After incubation, 1mL of the supernatant suspension was collected to measure the absorbance at 600 nm . Auto-aggregation (%) was calculated with the following equation:
Auto-aggregation (%) = (1- At / A0h) ×100
Where, At represents the absorbance at t = 1, 2, 3, 4 or 5 h and A0 is the absorbance at t = 0. The test was performed in triplicate
Co- aggregation assay
The co-aggregation potential of LAB isolates with different bacterial strains viz. Bacillus subtilis MTCC441, Bacillus cereus MTCC430, Salmonella enteric MTCC1166, Shigella flexneri MTCC1457, Streptococcus pneumonia MTCC4673, Staphylococcus aureus MTCC7443, Staphylococcus epidermidis MTCC435, Pseudomonas aeruginosa MTCC4673, E. coli MTCC40, Clostridium perfringens MTCC 450, Listeria monocytogens MTCC 657 and yeast Candida albicans MTCC1637, was examined. LAB and the test pathogenic bacteria were grown in MRS broth and nutrient broth, respectively, for 24h at 35±1°C. Bacterial suspension (108 cfu/ml) were formulated as described above as in the auto- aggregation in above method, equal volume of LAB and pathogenic strains (1:1 v/v) were mixed and incubated at 35±1°C without agitation (Prabhurajeshwar et al., 2017). Absorbance, A600 of the mixture represent above, was supervised during incubation at 4 h, percentage of co- aggregation was calculated using the following equation:
Co-aggregation (%) = [(Apathogen +A LAB)] / 2-Amix ( Apathogen +ALAB) /2×100
Cell surface hydrophobicity: The bacterial adhesion to hydrocarbons assay was performed according to the method Xu et al., (2009) with slightly modification to determine the cell surface hydrophobicity. Bacterial cells were suspended in phosphate buffered saline (PBS) pH 7.2 to 108 cfu /ml. Then, equal proportions of viable bacterial suspension and solvent (Xylene) were mixed by vortexing and incubated at 35±1°C for 10 min for temperature equilibration. To separate the mixture into two phases, the mixture was again vortexed briefly and allowed to stand for 5min. the aqueous phase was removed and its absorbance was measured at 600 nm. The results were reported as percentages according to the formula: H% = [(Ao – A) / Ao] ×100
Where, Ao and A are absorbance before and after mixing with xylene, respectively.
Data analysis
All measurements of antagonistic activity, Auto-aggregation assay, Co- aggregation assay and Cell surface hydrophobicity were performed in triplicate. The data was expressed in the mean and standard deviation (± S.D) in triplicates.
Molecular Identification
The best selected isolates on the basis of their Auto-aggregation assay, Co- aggregation assay, Cell surface hydrophobicity and antagonistic activity against all the pathogens, was identified using 16s rRNA by Sanger dideoxy sequencing. The purified culture was sent for commercial sequencing. The sequence data obtained was compared using BLAST tool (Basic Local Alignment Search Tool).
Sequencing the DNA
The sequence data obtained was deposited to NCBI database with BLAST analysis for molecular identification.
RESULTS AND DISCUSSIONS
Total 8 isolates of LAB were isolated from curd , cow milk, buttermilk, goat milk and paneer samples based on their morphological, physiological and biochemical tests (Abdullah and Osman, 2010; Bisen et al., 2013; Guetouache and Guessas et al.,2015 and Kang et al., 2019) as per the guidelines of Bergey’s Manual of Systematic Bacteriology (Hammes et al., 2009). All the 8 isolates were tested for antagonistic effect against common enteric pathogens Bacillus subtilis MTCC441, Bacillus cereus MTCC430, Salmonella enteric MTCC1166, Shigella flexneri MTCC1457, Streptococcus pneumonia MTCC4673, Staphylococcus aureus MTCC7443, Staphylococcus epidermidis MTCC435, Pseudomonas aeruginosa MTCC4673, E. coli MTCC40, Clostridium perfringens MTCC 450, Listeria monocytogens MTCC 657 and yeast Candida albicans MTCC1637 (Table1). The antagonistic activity of LAB is due to the production of antimicrobial compounds such as lactic acid, ethanol, hydrogen peroxide (H2O2), diacetyl, reuterin, bacteriocin and biosurfactants (Cizeikiene et al., 2013; Guetouache and Guessas et al., 2015, Gaspar et al., 2018). Amongst all the 8 isolates only C9 showed antagonistic spectrum against all the pathogens (Table 1). All the 8 isolates shows antagonistic spectrum against Staphylococcus aureus, Staphylococcus epidermidis and Salmonella enteric. Except c20 all the species shows maximum zone of inhibition against Bacillus subtilis, highest by C9 (19mm), c12 (18mm), G4 (17mm) followed by P37, B23, b1 and least by C28 (Table 1; Figure 2). Except c12 and B23, All the LAB isolates showed antagonistic spectrum against Bacillus cereus (Figure 1(B)), similar results were reported by Nigam et al., 2012. Except b1, all the isolates shows highest zone of inhibition against Streptococcus pneumonia by C9 (22mm), followed by C28 (18mm), G4 (16mm), B23 (12mm), c12 (11mm) and least by P37 (10mm). Against Pseudomonas aeruginosa highest antagonistic spectrum by C9 (20mm) followed by C28 and P37 (17mm), G4 (17mm), c12 (13mm) and least by c20 (12mm) (Table1; Figure 1(A)). Except C28 and P37 all showed antagonistic acitivity against E.coli Highest by C9 (22mm) and lowest by B23 (10mm). Except G4 all isolates showed zone of inhibition against Salmonella enteric. Against Candida albicans, highest zone of inhibition showed by C9 (19mm), C28 (17mm), G4 (15mm) followed by c20, c12 and lowest by P37 and b1 (11mm). Except P37, all showed maximum zone of inhibition against Clostridium perfringens. Highest zone of inhibition against Listeria monocytogens shown by C9 (23mm), G4 (20mm), C28 and B23 (19mm) followed by P37, C28 and c12. All the 8 isolates of LAB can be used as alternative in food preservatives and also replace chemical additives (Costa et al., 2018 and Hu et al., 2017).
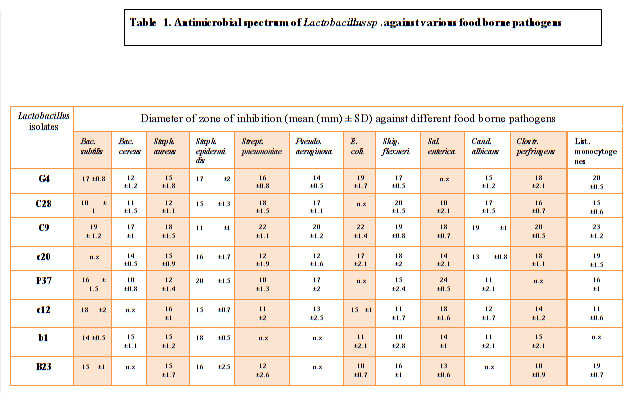 |
Table 1. Antimicrobial spectrum of Lactobacillus sp .against various food borne pathogens |
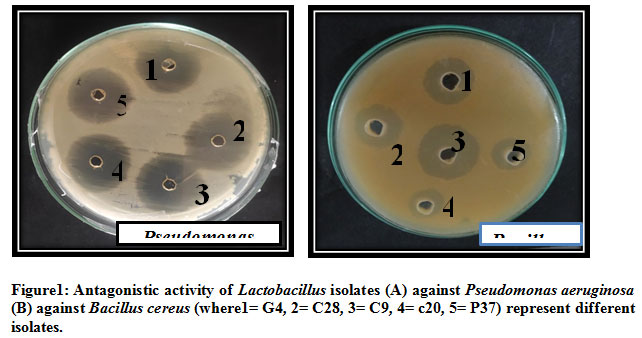 |
Figure 1: Antagonistic activity of Lactobacillus isolates (A) against Pseudomonas aeruginosa (B) against Bacillus cereus (where1= G4, 2= C28, 3= C9, 4= c20, 5= P37) represent different isolates |
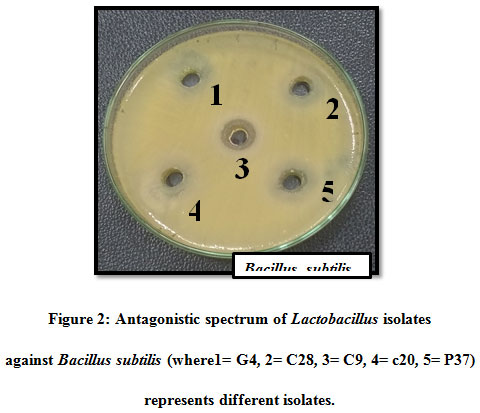 |
Figure 2: Antagonistic spectrum of Lactobacillus isolates against Bacillus subtilis (where1= G4, 2= C28, 3= C9, 4= c20, 5= P37) represents different isolates |
They also satisfy consumers’ demands for fresh, healthy and safe food (Ahmad et al., 2017). Cotter et al., 2013 stated that the use of bacteriocin as food preservative meet consumers’ demand for natural preservation and also considered additional safety measure to minimally processed products. Yang et al., 2014; Ahmad et al., 2017 and Kang et al., 2019 reported that bacteriocins were able to kill target microorganism by disrupting membrane integrity which little induce resistance, since their fragments do not interact also with target cells and the best potential solution for growing problems of microbial resistance to antibiotics. One of the most important prerequisite for the colonization of the host intestinal tract is the adhesion ability of probiotics to intestinal mucus and enterocytes has been also proposed as one of the important selection criteria for potential probiotics (Xu et al., 2009). Through in vitro model system adhesion ability of probiotics has been studied, which are commonly used in selection and assess of probiotics strains for in vivo studies (Baick and Kim, 2015).
Table 2. Mean auto-aggregation percentage of different isolates of Lactobacillus over a period of 5 h.
| STRAIN | Auto-aggregation (%) | ||||
| 1 h | 2 h | 3 h | 4 h | 5 h | |
| G4 | 11.3±1.3 | 36.6±1.4 | 61±1 | 70.6±1.6 | 75±1.1 |
| C28 | 14.6±1.3 | 32.3±1.7 | 44.3±3 | 61.3±1.2 | 76.3±0.1 |
| C9 | 26.6±2.4 | 54.3±1.4 | 67.3±0.1 | 83±2.1 | 91.6±1.2 |
| c20 | 13.3±1.2 | 20±0.8 | 36.6±1.1 | 50±1.7 | 66±0.8 |
| P37 | 16.3±1.2 | 27.6±2 | 44.6±1.4 | 52±1.1 | 61.3±2.6 |
| c12 | 8±0.8 | 21.6±1.2 | 40±1 | 56.3±2 | 60.3±2 |
| b1 | 15.4±2.1 | 20±1.5 | 35±0.8 | 44.3±1.3 | 50.5±2.1 |
| B23 | 9.4±1.5 | 18.6±2 | 45.6±1.1 | 56.3±1.2 | 64.2±1.3 |
All values are mean ± standard deviation of triplicate
For in vitro adhesion tests generally immobilized intestinal mucus and human enterocyte-like Caco3 cell cultures are mostly used but these methods are expensive and time consuming, (Andrabi et al., 2016) therefore, an auto- aggregation test together with cell surface hydrophobicity (CSH) have been extensively used for the preliminary screening and identification of potential adherent bacteria with suitable commercial (Zielińska et al., 2015). Bajaj et al., (2015) stated that Cell surface hydrophobicity by xylene partition has been used as an indirect test to estimate the adhere ability of probiotics to epithelial cells. The results of auto- aggregation assay are shown in table 2. C9 showed highest auto-aggregation ability of 91.6% followed by C28 (76.3%), G4 (75%), c20 (66%), B23 (64.2%), P37 (61.3%), c12 (60.3%) and least by b1 (50.5%). Prabhurajeshwar et al., (2017) stated that the auto- aggregation % increased with increased in incubation time and results were persistent.
Baick and Kim, (2015) also reported 90% as the highest auto- aggregation % similarly Andrabi et al., 2016 documented 94.97 highest auto aggregation % at 37ºC and at 25 ºC 96.54% was documented as the highest auto-aggregation percentage. All the eight isolates were further tested for their co-aggregation ability with different pathogens viz., Bacillus subtilis MTCC441, Bacillus cereus MTCC430, Salmonella enteric MTCC1166, Shigella flexneri MTCC1457, Streptococcus pneumonia MTCC4673, Staphylococcus aureus MTCC7443, Staphylococcus epidermidis MTCC435, Pseudomonas aeruginosa MTCC4673, E. coli MTCC40, Clostridium perfringens MTCC 450, Listeria monocytogens MTCC 657 and yeast Candida albicans MTCC1637 (Table 3). Excellent co-aggregation potential was exhibited by all the isolates except some isolates showed low co-aggregation like G4 against Salmonella enteric i.e 34.1%, c20 against Bacillus cereus (33.3%), P37 and c12 against E. coli (36.6% and 32.5%), b1 against Pseudomonas aeruginosa and Listeria monocytogens (30.5% and 23.2%) and B23 against Streptococcus pneumonia and Pseudomonas aeruginosa however, with other pathogens it showed relatively batter co-aggregation (Table 3).
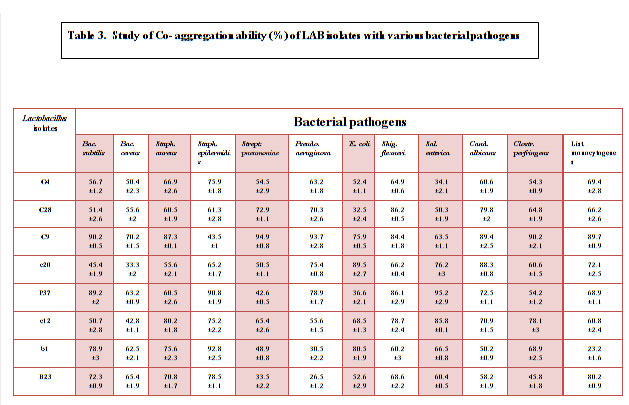 |
Table 3: Study of Co- aggregation ability (%) of LAB isolates with various bacterial pathogens |
Overall all the isolates had remarkable co-aggregation potential with pathogens. Isolates tested in this study showed a variable hydrophobicity with values ranging from 0.5% to 79% (Table 4). Amongst all the eight isolates C9 showed noticeable higher hydrophobicity. Among the eight isolates, isolate C9 isolated from cow milk exhibit antagonistic activity against all the food borne pathogens and common enteric pathogens. Analysis of the 16S rDNA sequence of isolate C9 showed high homology with that of available strains of Lactobacillus fermentum (National Center for Biotechnology Information, NCBI database); thus, isolate C9 was identified as Lactobacillus fermentum and designated as Lactobacillus fermentum C9. The sequence has been submitted to GenBank under accession number MN421922.
Table 4. Cell surface hydrophobicity
| Lactobacillus isolates | Hydrophobicity (%) |
| G4 | 0.5±1.2 |
| C28 | 51.4±1.2 |
| C9 | 79.0±0.7 |
| c20 | 5.2±0.5 |
| P37 | 8.0±2.2 |
| c12 | 37.1±2.5 |
| b1 | 15.6±1.8 |
| B23 | 19.1±0.2 |
CONCLUSION
Thus, it may be safely concluded that all eight isolates of Lactobacillus isolated in this study can be safely used as food preservatives.
References
Abdullah S A and Osman M M (2010). Isolation and identification of lactic acid bacteria from raw cow milk, white cheese and Rob in Sudan. Pak. J. Nutr. 9: 1203-1206.
Adeyemo S M, Agun T F and Ogunlusi E D (2018). Antimicrobial activity of lactic acid bacteria isolated from ‘Pupuru’: An African fermented staple against food borne- pathogens. J. Mol. Bio. Biotech. 3:1-5.
Ahmad V, Khan S M, Jamal S M Q, Alzohairy A, Karaawi Al A M and Siddiqui U M (2017). Antimicrobial potential of bacteriocins: in therapy, agriculture and food preservation. Int. J. Antimicro. Agents. 49: pp1-11.
Akkoc N, Ghamat A and Akcelik M (2011). Optimisation of bacteriocin production of Lactococcus lactis subsp. lactis MA23, a strain isolated from Boza. Int J. of Dairy Tech. 64;3.
Andersson D I and Hughes D (2010). Antibiotic resistance and its cost: is it possible to reverse resistance? Nat Rev Microbiol. 8(4):260–71.
Andrabi T S, Bhat B, Gupta M and Bajaj K B (2016). Phytase producing potential and other functional attributes of lactic acid bacteria isolates for prospective probiotics applications. Probiotics & Antimicro. Prot. 8; 121-129.
Astó E, Méndez L , Audivert S, Codina F A and Espadaler J (2019). The Efficacy of Probiotics, Prebiotic Inulin-Type Fructans, and Synbiotics in Human Ulcerative Colitis: A Systematic Review and Meta-Analysis. Nutrients .11, 293.
Bah A, Albano H, Barbosa B J, Fhoula I, Gharbi Y, Najjari A, Boudabous A, Teixeira P and Ouzari I H (2019). Inhibitory effect of Lactobacillus plantarum FL14 against Food borne pathogens in artificially contaminated fermented tomato juices. BioMed Res. Inter.
Bajaj B K, Razdan K, Claes I J I and Lebeer S (2015). Probiotics attributes of the newly isolated lactic acid bacteria from infants’ gut. J. Microbiol. Biotechnol. Food Sci.5; 109-115.
Bisen S P, Sharma R, Sanodiya S B, Thakur S G, Jaiswal P, Pal S and Sharma A (2013). Characterization of Lactic acid bacteria from raw milk samples of goat, sheep, camel and buffalo with special elucidation to lactic acid production. Brit Micro Res J. 3(4):743-752.
Campana R, Hemert V S and Baffone W (2017). Strain‑specific probiotic properties of lactic acid bacteria and their interference with human intestinal pathogens invasion. Gut Pathog. 9:12.
Centers for Disease Control and Prevention (CDC). Food borne disease active surveillance network (FoodNet): Food Net surveillance report for 2012 (final report). Atlanta: CDC; 2014.
Centers for Disease Control and Prevention (CDC). Incidence and trends of infection with pathogens transmitted commonly through food- food borne disease active surveillance network, 10 U.S. sites, 1996-2012. MMWR Morb Mortal Wkly Rep.2013; 62(15): 283-7.
Chiang S S and Pan T M (2011). Beneficial effects of Lactobacillus paracasei paracasei NTU 101 and its fermented products. Appl. Microbio. Biotech. 93(3), 903–916.
Cizeikiene D, Juodeikiene G, Paskevicius A, Bartkiene E (2013).Antimicrobial activity of lactic acid bacteria against pathogenic and spoilage microorganisms isolated from food and their control in wheat bread. Food Control 31, 539–545.
Colomer V M, Faolony G, Tigchelaar F E, Wang J, Tito Y R, Schiweck C, Kurilshikov A, Joossens M, Wijmenga C, Claes S, Oudenhove V L, Zhernakova A, Silva V S and Raes J ( 2019). The neuroactive potential of the human gut microbiota in quality of life and depression. Nature Mic.4, 623-632.
Costa D J R, Voloski S L F, Rafael G, Mondadori G R, Duval H E and Fiorentini M A(2019). Preservation of Meat Products with Bacteriocins Produced by Lactic Acid Bacteria Isolated from Meat. Food Qual.12
Cotter P D, Ross R P and Hill C (2013). Bacteriocins—a viable alternative to antibiotics? Rev. Microbiol. 11, 95–105
Cotter PD, Hill C and Ross R P (2005). Bacteriocins: developing innate immunity for food. Nat. Rev. Microbial.3: 777-778.
Coventry M J, Gordon J B, Wilcock A, Harmark K, Davidson B E, Hickey M W, Hillier A J and Wan J (1997). Detection of bacteriocins of lactic acid bacteria isolated from foods and comparison with pediocin and nisin. J. Appl. Microbiol. 83 (2); 248-58.
Cross M L (2002). Microbes versus microbes: immune signals generated by probiotic lactobacilli and their role in protection against microbial pathogens. FEMS Immunol. Med. Microbiol. 34; 245-253.
Dhanasekaran D, Subhasish Saha N, Thajuddin M, Rajalakshmi M and Panneerselvam A (2010). Probiotic effect of Lactobacillus isolates against bacterial pathogens in fresh water fish. J. Coast. Dev. 13(2): 103–112.
Dickinson B and Surawicz C M (2014). Infectious diarrhea: an overview. Curr Gastroenterol Rep.16(8):399.
Feliatra F, Muchlisin A Z, Teruna Y H, Utamy R W, Nursyirwan N and Dahliaty A (2018). Potential of bacteriocins produced by probiotics bacteria isolated from tiger shrimp and prawns as antibacterial to Vibrio, Pseudoonas and Aeromonas species on fish. F1000Research.7:415.
Fuller R (1989). A review: Probiotics in man and animals. J. Appl. Bact. 66; 365-378.
Gao Z,Daliri B M E, Wang J, Liu D, Chen S, Ye X and Ding T (2019). Inhibitory Effect of Lactic Acid Bacteria on Foodborne Pathogens: A Review. J. Food Prot. 82 (3). 441-453.
Gaspar C, Donders G G, Oliveira De P R, Queiroz A J, Tomaz C, Oliveira De M J and Oliveira De P A (2018). Bacteriocin production of the probiotic Lactobacillus acidophilus KS400. AMB Expr.8:153.
Guetouache M and Guessas B (2015). Characterization and identification of lactic acid bacteria isolated from traditional cheese (Klila) prepared from cow’s milk. Afr. J. Microbiol. Res. 9(2), pp. 71-77.
Guo L, He M Y, Li M, Chen W D, Chen X, Huang J Y and Gu X R (2017). Inhibitory effects of single and mixed lactic acid bacteria on intestinal pathogens. Food Sci. Technol. 42: 26-30.
Hammes P H and Hertel C, Lactobacillaceae. In: Vos P.D., Garrity, G.M., Jones, D., Krieg, N.R., Ludwig, W., Rainey, F.A., Schieifer, K.H., Whitman, W.B. Ed, Bergey’s Manual of Systematic Bacteriology, 3rd Springer, New York, (2009) pp 456-479.
Hu Y, Liu X and Shan C (2017). Noval bacteriocin produced by Lactobacillus alimentarius FM-MM 4 from a traditional Chinese fermented meat nanx wudl: purification, identification and antimicrobial characteristics. Food Control .77: 290-297.
Hwanhlem N, Chobert J.-M and H-Kittikun A (2014). Bacteriocin-producing lactic acid bacteria isolated from mangrove forests in southern Thailand as potential bio-control agents in food: Isolation, screening and optimization. Food Control. 41; 202–211.
Jolly L,Vincent F J S, Duboc P, Neeser R J (2002). Exploiting exopolysaccharides from lactic acid bacteria. Antonie van Leeuwenhoek. 82; 367–374.
Kang H J and Im S H (2015). Probiotics as an immune modulator. J. Nutr. Sci. Vitaminol. 61: 103-5.
Kermanshahi K R and Qamsari M E (2015). Inhibition effect of lactic acid bacteria against food born pathogen, Listeria monocytogenes. Food Biotechnol.2(4): 11-19.
Kumari A, Garg P A, Makeen K and Lal M (2008). A bacteriocin production on soya nutri nuggets extract medium by Lactococcus lactis Lactis CCSUB202.Int. J. dairy Sci. 3(1).49-54.
Li S J, Gao X M, Cui B C, Liu C C, Mou H, Zhao X and Wang D Z (2014). Effects of irradiation on antimicrobial activity of weak organic acid preservatives. Food Sci. 35: 58-62.
Masuda Y, Zendo T and Sonomoto K (2012). New type of non-lantibiotic bacteriocins: circular and leaderless bacteriocins. Beneficial Microbes.3:3–12.
Mejlholm O and Dalgaard P (2015). Modelling and predicting the simultaneous growth of Listeria monocytogenes and psychrotolerant lactic acid bacteria in processed seafood and mayonnaise- based seafood salads. Food Microbiol.46: 1-14.
Millette M, Dupont C, Archambault D and Lacroix M (2007). Partial characterization of bacteriocins produced by human Lactococcus lactis and Pediococcus acidilactici J. Appl. Microbiol. 102; 274-282.
Molina V, Médici M, Font de Valdez G and Taranto M P (2012). Soybean-based functional food with vitamin B12-producing lactic acid bacteria. J. Functional Foods. 4(4); 831–836.
Nigam A, Kumar A, HV M and Bhola N (2012). In-vitro Screening of antibacterial activity of lactic acid bacteria against common enteric pathogens. J Biomed Sci. 1.4:2.
Oroojalian F, Kasra Kermanshahi R, Azizi M and Bassami M (2010). Phytochemical composition of the essential oils from three Apiaceae species and their antibacterial effects on food-borne pathogens. Food Chem. 120(3): 765-770.
Østergaard N B, Eklöw A and Dalgaard P (2014). Modelling the effect of lactic acid bacteria from starter and aroma culture on growth of Listeria monocytogenes in cottage cheese. Int. Food Microbiol. 188:15-25.
Prabhurajeshwar C and Chandrakanth K R (2017). Probiotic potential of Lacobacilli with antagonistic activity against pathogenic starins: An in vitro validation for the production of inhibitory substances. Biomed. J. 270-283.
Ramos C L, Thorsen L, Schwan R F and Jespersen L (2013). Strain specific probiotics properties of Lactobacillus fermentum, Lactobacillus plantarum and Lactobacillus brevis isolates from Brazilian food products. Food Microbiol. 36:22-29.
Ren C, Du H and Xu Y (2017). Advances in microbiome study of traditional Chinese fermented foods. Acta Microbiol. Sin. 57: 885– 898.
Sari M, Suryanto D and Yurnaliza (2018). Antimicrobial activity of lactic acid bacteria isolated from bekasam against Staphylococcus aureus ATCC 25923, Escherichia coli ATCC 25922 and Salmonella sp.IOP Conf. Series: Earth and Environ. Sci.130
:012011.
Sharma C, Singh P B, Thakur N, Gulati S, Gupta S, Mishra K S and Panwar H (2017). Antibacterial effects of Lactobacillus isolates of curd and human milk origin against food-borne and human pathogens. 3 Biotech. 7: 31– 39.
Shimizu M, Hashiguchi M, Shiga T, Tamura H and Mochizuki M (2015). Meta-Analysis: Effects of Probiotic Supplementation on Lipid Profiles in Normal to Mildly Hypercholesterolemic Individuals. Plos One. 10 (10).
Sieladie D, Zambou N, Kaktcham P(2011). Probiotic properties of lactobacilli strains isolated from raw cow milk in the western highlands of Cameroon. Innovative Romanian Food Biotechnology.9:12–28.
Simova D E, Beshkova B D and Dimitrov P Z (2008). Characterization and antimicrobial spectrum of bacteriocins produced by lactic acid bacteria isolated from traditional Bulgarian dairy products. J. Appl. Microbio.106:692-701.
Tankoano A, Diop B M, Lingani S H, Mbengue M, Kaboré D, Traoré Y and Savadogo A (2019). Isolation and Characterization of Lactic Acid Bacteria Producing Bacteriocin like Inhibitory Substance (BLIS) from “Gappal”, a Dairy Product from Burkina Faso. Adva. Microbi. 9, 343-358.
Tesfaye A (2014). Antagonism and primary in vitro probiotic evaluation of lactic acid bacteria recovered from Ergo. J Agr Biol Sci.7: 240-245.
Wang H, Wei X C, Min L and Zhu Y L (2018). Good and bad: gut bacteria in human health and disease. Review; Medi. Biotech. 32,5, 1075- 1080.
Wemmenhove E, Valenberg Van F J H, Hooijdonk Van M C A, Wells-Bennik J H M and Zwietering H M (2017). Factors that inhibit growth of Listeria monocytogenes in nature-ripened Gouda cheese: a major for undissociated lactic acid. Food Control. 84: 413-418.
Xu H, Jeong H S, Lee H Y and Ahm J (2009). Assessment of cell surface properties and adhesion potential of selected probiotics strains. Lett Microbiol. 49,434-442.
Yang S C, Lin H C, Sung T C and Fang Y J (2014). Antibacterial activities of bacteriocins: application in foods and pharmaceuticals. Front Microbiol. 5;241.
Yang E, Fan L, Jiang Y, Doucette C and Fillmore S (2012). Antimicrobial activity of bacteriocin producing lactic acid bacteria isolated from cheeses and yogurts. AMB Express. 2:48.
Yang Y, Latorre D J, Khatri B, Kown M Y, Kong W B, Teague D K, Graham E L, Wolfenden D A, Mahaffey D B, Baxter M, Velasco H X, Guzman M R, Hargis M B and Tellez G (2017). Characterization and evaluation of lactic acid bacteria candidates for intestinal epithelial permeability and Salmonella typhimurium colonization in neonatal turkey poults. Poult. Sci. 97:515-521.
Zhang J S, Wu S S, Zhao H L, Ma L Q, Li X, Ni Y M, Zhou T and Zhu L H (2018). Culture –dependent and independent analysis of bacterial community structure in Jiangshui, a traditional Chinese fermented vegetable food. LWT Food Sci. Technol. 96: 244-250.
Zhang J, Tian G Z, Wang H J and Wang R A (2011). Advances in antimicrobial molecular mechanism of organic acids. Chin. J. Anim. Vet. Sci. 42: 323-328.
Zielińska D, Rzepkowska A, Radawska A and Zieliński K (2015). In vitro screening of selected probiotics properties of Lactobacillus strain isolated from traditional fermented cabbage and cucumber. Curr Microbiol.70; 183-194.


