1Department of Microbiology, Faculty of Science, Annamalai University,
Chidambaram, Tamil Nadu, India
2Department of Microbiology, Mother Theresa Post Graduate & Research
Institute of Health Sciences, Puducherry, India
Corresponding author email: drsumathimicro@gmail.com
Article Publishing History
Received: 09/10/2021
Accepted After Revision: 12/11/2021
In an array of identifying safe antimicrobial compounds, bacteriocin producing Lactobacillus strain have been investigated in this study from the daily consuming food resources of humans. Till now, the best studied bacteriocins are nisin A produced by Lactobacillus lactis and pedocin PA-1 synthesized by Pediococcus acidilactici which have been accredited as a preservative in the food industries by the World Health Organization (WHO). For this study, four different milk and dairy products viz., curd, cheese, yoghurt and butter were collected from the local markets of Karaikal region, Puducherry, India and were used for the isolation of Lactobacillus species using MRS agar. Totally, five morphologically distinct strains were collected and were initially named as MPD 1 to MPD 5. During the screening process of bacteriocin production, the strain MPD 5 showed maximum antimicrobial activity against Vibrio cholerae MTCC 3906 with 900AU/ml.
This strain was molecular identified as Lactiplantibacillus plantarum MDP 5 based on 16S rRNA partial sequence method. This 16S rRNA partial sequence was submitted to the NCBI nucleotide GenBank and provided with the accession number, MW301154.1. Further, this strain revealed an enhanced production of bacteriocin using the standardized physicochemical factors such as pH 7, 35°C, 2% fructose and 1% peptone. Furthermore, these optimal conditions revealed more than 2-fold increase in the bacteriocin production. All the above information suggesting the possibilities of bacteriocin for the bioindustrial production using the L. plantarum MDP 5 of this study and its future prospects for the investigation of biocidal activities against many highly infectious pathogens of human and veterinary.
Antimicrobial, Bacteriocin, Growth Kinetics, Lactiplantibacillus Plantarum,
Milk And Dairy Products, Optimized Production.
Punithavalli T, Prince C.P, Sumathi V.Screening and Evaluation of a Potential Bacteriocin-Producing Lactobacillus Species in Milk and Dairy Products for their Enhanced Production Collected from Puducherry, India. Biosc.Biotech.Res.Comm. 2021;14(4).
Punithavalli T, Prince C.P, Sumathi V.Screening and Evaluation of a Potential Bacteriocin-Producing Lactobacillus Species in Milk and Dairy Products for their Enhanced Production Collected from Puducherry, India. Biosc.Biotech.Res.Comm. 2021;14(4). Available from: <a href=”https://bit.ly/3n8GJ1g“>https://bit.ly/3n8GJ1g</a>
Copyright © This is an Open Access Article distributed under the Terms of the Creative Commons Attribution License (CC-BY) https://creativecommons.org/licenses/by/4.0/, which permits unrestricted use distribution and reproduction in any medium, sources the original author and sources are credited.
INTRODUCTION
Lactobacillus species are common for the application of fermentation as well as preservation of broad class of milk, vegetable, meat products, etc. There are antimicrobial compounds derived from the Lactobacillus strains viz., diacetyl, hydrogen peroxide, organic acids as well as bacteriocins which reveal a key role in assuring the safety and lengthening the shelf-life of many food products. Since the last 20 years, antimicrobial significance of Lactobacillus strains has been globally studied (Daeschel 1989; Jack et al. 1995).
Increasing number of consumers demanding the organic based bioactive products in the recent years enhance the interest of applications using naturally derived inhibitory substances which could alternate the chemical counterpart. Among such natural bioactive compounds, bacteriocin could be in the leading interest which attract many researchers globally (Heredia-Castro et al. 2015; Balan et al. 2019a; Balan et al. 2019b).
Synthesis of bacteriocin is a common phenomenon among Lactobacillus species. Bacteriocins are bioactive proteins which usually act antagonistic towards Gram-positive bacteria or closely related to the synthesizer microbe and the appreciable bioactivities were recorded against Staphylococcus aureus, Clostridium tyrobutyricum, Listeria innocua, Listeria monocytogenes and Bacillus cereus (Klaenhammer 1988). Further, bacteriocin rarely evidenced its bioactivities against gram negative bacteria such as Helicobacter pylori Salmonella typhimurium, Campylobacter jejuni and Escherichia coli (Rammelsberg and Radler 1990; Reis et al. 2012). Bacteriocin production have also been reported from Leuconostoc, Carnobacterium, Pediococcus, and Lactococcus species (Kohoutova et al. 2020; Nebbia et al. 2021).
The best studied bacteriocins are nisin A produced by a L. lactis and pedocin PA-1 synthesized by a Pediococcus acidilactici which have been accredited as a preservative in the food industries by the World Health Organization (WHO) (Biscola et al. 2013). Bacteriocins have many positive features which compel it special for numerous different applications. Bacteriocin from Lactobacillus are often having extreme thermal and pH tolerance. Further, these peptides are also known for their colourless, odourless, and tasteless, which further enhance their added advantage for many potential usefulness (Negash and Tsehai 2020).
In the present investigation, fermented food from the local markets of Karaikal, Puducherry have been aimed for the isolation and screening of bacteriocin producing Lactobacillus strains which explore the significance of indigenous microbiota. Indigenous microbes of various natural or geographical habitats have unique biochemical, functional as well as metabolic properties (Balan et al. 2013a; Balan et al. 2013b; Ahmad et al. 2020; Chen et al. 2021). Hence, this study concentrated on native species and the potential isolate have further been characterized for its enhanced bacteriocin production.
MATERIAL AND METHODS
Sampling and isolation of Lactobacillus strains was performed using four different milk and dairy products viz., curd, cheese, yoghurt and butter were collected from the local markets of Karaikal region, Puducherry, India. All the samples were individually processed for the isolation of Lactobacillus strains. One gram of sample was serially diluted with pre-sterilised bacteriological saline water and spread plated on MRS agar medium (HiMedia, Cat. No. M641) plates. After two days incubation at 37°C, morphologically distinct colonies were axenic cultured in newly prepared MRS agar plates. The purity of the bacterium was microscopically confirmed after Gram’s staining method (Harley 2013). All the axenic strains were stored under lyophilized conditions for further use.
Screening of bacteriocin producing bacteria was done using an indicator pathogenic bacterium, Vibrio cholerae MTCC 3906 purchased from the Microbial Type Culture Collection, CSIR-Institute of Microbial Technology, Chandigarh, India. The strain was overnight cultured at 37°C on tryptone soya broth (HiMedia, Cat. No. LQ508) and the OD of the culture was tuned to an inoculum concentration of 105 CFU/ml for all the bioactivity studies. The quantitative bacteriocin production was determined by microtiter well-plate assay as earlier described in the previous studies (Kang and Lee 2005). The bioassay was performed in 96-well flat bottom polystyrene microtitre plates with lids (Tarsons, India). In this assay, well plates were filled with 100μL of serially diluted bacteriocin containing cell free supernatant followed by the addition of 20μL indicator strain, V. cholerae MTCC 3906 at a concentration of 105 CFU/ml assessed using optical density at 620nm and 80μL tryptone soy broth. The growth control well plate has 100μL of phosphate buffer (pH 7), 20μL of indicator strain and 80μL tryptone soya broth.
The susceptibility control well plate has 100μL of 4mg/ml streptomycin containing phosphate buffer (pH 7), 20μL of indicator strain and 80μL of tryptone soy broth. The well plates were incubated for 6hrs at 37°C, further, the absorbance was estimated at 620nm with a microplate reader (Biotek Elx808, WI, USA). The growth inhibition percentage were estimated as given below: Percentage growth inhibition = [(1 − (As/Ac)] × 100, where As denotes the absorbance of wells having the test samples and Ac denotes the absorbance of wells without any added bioactive sample (control). Bacteriocin activity was represented here with the arbitrary unit (AU) in which the AU is defined as reciprocal of the maximum dilution of sample forming 50% growth inhibition of indicator strain.
The study of extracellular or intracellular bacteriocin production was performed with the individual isolates. The strains were cultured in 30ml screw cap tubes containing aliquot MRS broth (HiMedia, Cat. No. M369). After two days incubation at 37°C, centrifugation was performed on the cultured broths for the separation of cell pellet and cell free supernatant at 3000rpm 15 min. The separated cell pellet dissolved in 50ml phosphate buffer and sonicated at 20KHz for 45 sec. using Ultra-sonicator (Hielscher, USA). The same centrifugation condition was applied to remove the cell debris formed during the sonication process and the supernatant was used for the estimation of bacteriocin production whereas the cell free supernatant collected from cultured broth was instantly applied for the bacteriocin bioassay. This experiment revealed whether the isolate produced extracellular or intracellular bacteriocin or observed from both locations.
Molecular identification of potential bacteriocin producer was done using 16S rRNA partial sequence using the Eubac set of primers, 27F (5’-AGAG TTTG ATCM TGGC TCAG-3’) and 1492R (5’-GGTT ACCT TGTT ACGA CTT-3’). The phylogenetic tree was drawn with neighbour joining procedure (Saitou and Nei 1987) against the maximum similar sequences available in the NCBI nucleotide GenBank database using MEGA 7 software (Kumar et al. 2016). Optimization of enhanced bacteriocin production was performed in this study in which the potential strain was standardized for the possible enhanced production by adopting search technique varying one condition at a time and the standardized conditions were fixed for subsequent estimations.
The optimization of bacteriocin production was studied in 250ml conical flask with 100ml working volume of yeast glucose broth [10] has the following composition of yeast extract (10g/L) and glucose (20g/L) with the final pH of 7.5 ± 0.2 along with the subsequent conditions of 37°C temperature, agitation at 150rpm and one ml inoculum. The inoculum preparation and bacteriocin quantification was performed using the same applied procedure as mentioned in the screening of bacteriocin producing bacteria.
Growth kinetics profile as a function of time on bacteriocin production was carried out with the potential strain. It was studied for the required time of maximum bacteriocin production with reference to the cell biomass concentration at 12hrs regular time intervals from 0hrs to 144hrs. The estimations were performed with a portion of cultured broth next by the separation of cell biomass and cell free supernatant with the help of centrifugation at 3000rpm for 15 min. Bacterial cell growth was estimated from the dry weight of cell biomass resulted from the hot air oven dried centrifuged cell pellets at 50°C for 30min. and bacteriocin quantitative estimation was directly determined from the cell free supernatant.
Recapitulation of different biotic and abiotic growth parameters was performed in this study. Optimizing parameters like pH, temperature, carbon and nitrogen take part a significant role in the enhanced bacteriocin production. Influence of pH conditions from pH 5 to 9 were studied for the effective medium optimization, similarly, various temperature conditions from 20 to 50°C were investigated in the production medium for the enhanced production. Similarly, various carbon sources like fructose, galactose, sucrose, maltose, cellulose and starch were utilized at 2% concentration in the production medium as well as various nitrogen substrates like ammonium nitrate, malt extract, tryptone, beef extract and peptone were used at 1% concentration.
RESULTS AND DISCUSSION
Lactobacillus were generally regarded as safe (GRAS) microorganisms which playing a key role in the fermentation of foods. Lactobacillus strains were known to produce various antimicrobial compounds such as bacteriocins, D-isomers of amino acids, reuterin, hydrogen peroxide, acetaldehyde, CO2, and diacetyl (Cintas et al. 2001). Bacteriocins were ribosomally produced antimicrobial compounds which were active against bacterium of same species or other genera. Bacteriocins have been reported from both gram positive and negative bacteria. Nowadays, bacteriocin producing Lactobacillus have gained attention because of their non-toxic status and its uses as potential food preservative (Diop et al. 2007; Nebbia et al. 2021).
In this study four different fermentation products of milk and dairy such as curd, cheese, yoghurt and butter were taken for the isolation of Lactobacillus strains using MRS medium. Initially, the samples were serially diluted, spread plated on MRS agar medium and incubation for two days. There were five morphologically distinct strains obtained as axenic culture and these strains were initially referred using the voucher names viz., MPD 1 to MPD 5 in which the MDP represents the abbreviation of Milk and Dairy Products followed by serial number of the bacterial strains. All the strains were individually MRS broth cultured using screw cap tubes and screened for the intracellular and extracellular production of bacteriocin (Fig. 1).
Figure 1: Axenic strains cultured in 30ml screw cap tube containing
MRS broth for the screening of bacteriocin production

Among the isolates, only two strains revealed bacteriocin production viz., MDP 2 and MDP 5 through its antimicrobial activity against V. cholerae MTCC 3906 in which MDP 2 showed intracellular production and MDP 5 showed extracellular production and the rest of the strains revealed negligible bacteriocin activities. Further, the strain MDP 5 evidenced maximum bacteriocin activity of 900AU/ml followed by the strain MDP 2 with 150AU/ml. Based on the appreciable production of bacteriocin, the strain MDP 5 was chosen for molecular identification and enhanced bacteriocin production studies.
Similar to this study, an appreciable bacteriocin producing L. plantarum ST664BZ procured from a traditional cereal beverage of Bulgaria (Todorov and Dicks 2006). Likewise, bacteriocin purified from Enterococcus faecalis and E. hirae of homemade milk and dairy products revealed strong growth inhibitory activity against many foods borne pathogens (Sonbol et al. 2020).
The molecular identification of the isolated potential bacteriocin producing Lactobacillus strain, MDP 5 of this investigation was carried out with the 16S rRNA partial gene sequence method. The 16S rRNA partial gene sequence region was amplified using a PCR and the total sequence length of the amplified product was 1501bp. Based on BLASTn homology search, the amplified region revealed 100% sequence similarity with an available nucleotide database of the NCBI GenBank sequence and it was a type strain, L. plantarum NRIC 1960 with the accession number AB362769.1. Based on this highest gene sequence homology, the isolate MDP 5 was identified as Lactiplantibacillus plantarum as well as the sequence was deposited in NCBI nucleotide GenBank and provided with the accession number MW301154.1.
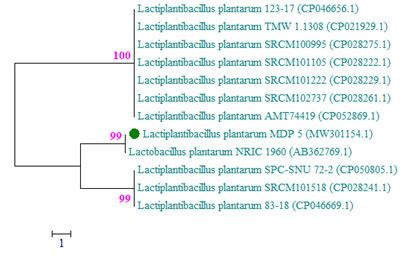
The genus Lactobacillus included in the family Lactobacillaceae and the phylum Firmicutes. The phylogenetic tree of L. plantarum MDP 5 was illustrated using the neighbour joining method with the twelve sequences of highest homology obtained from the NCBI GenBank nucleotide collection (Fig. 2). The exact molecular identification procedure was carried out for the identification of L. plantarum collected from the native fruits of Ecuadorian Amazon and a fermented drinks of China (Garzon et al. 2017; Pei et al. 2020).
Figure 2: The phylogenetic tree of L. plantarum MDP 5 based on evolutionary
analysis was studied using Neighbor-Joining method.

The growth kinetics profile of L. plantarum MDP 5 with regarding to the bacteriocin synthesis as a function of incubation time was examined (Fig. 3). The bacteriocin production was examined from the initial incubation period till the decline growth phase. Further, the peak production of bacteriocin was recorded during the end of exponential or beginning of stationary growth phase of the potential bacterium (60hrs) and it was maintained till the end of stationary or initiation of decline growth phase (96hrs) (Balan et al. 2019a; Balan et al. 2019b).
Figure 3: Growth kinetics profile as a function of time on bacteriocin
production using the L. plantarum MDP 5.
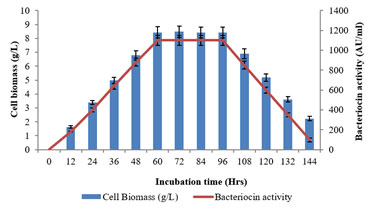
Furthermore, the highest production of bacteriocin, 1100±50AU/ml was achieved with the 8.43±0.42 cell biomass concentration. This growth dependant pattern of metabolite production revealed it is a primary metabolite (Balan 2014). The main difference between antibiotics and bacteriocins is that antibiotics were produced in the stationary growth phase and hold as secondary metabolites whereas bacteriocin synthesized during primary growth phase and act as primary metabolites (Beasley and Saris 2004; Balan et al. 2019a; Balan et al. 2019b).
Optimization of physicochemical parameters plays an important factor in the production process because it decides the commercialization value of any products (Sankar et al. 2013). Because each microbial strain has its own functional, metabolic and biochemical uniqueness, hence worth standardization of various physicochemical conditions requires to be done for the enhanced production of all the microbial based products (Mani et al. 2016; Balan et al. 2012; Balan et al. 2019a). Hence, this recapitulation study of pH, temperature, carbon and nitrogen parameters was carried out for the potential bacterium, L. plantarum MDP 5 of this study.
Figure 4: Effect of various pH conditions on the Bacteriocin
production using the L. plantarum MDP 5
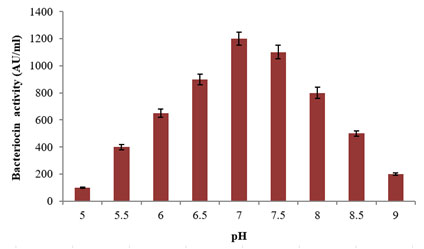
Among the tested pH conditions, the potential strain revealed maximum bacteriocin production at the pH 7 with 1200±50AU/ml activity followed by pH 7.5 and 6.5 with 1100±50AU/ml and 900±40AU/ml activities (Fig. 4). Further, the least bacteriocin activity of 100±5AU/ml was recorded at pH 5. This study proves the strong dependence of the L. plantarum MDP 5 on the hydrogen ion concentration of the surviving medium. A similar study was performed on temperature at varied conditions, the maximum bacteriocin production was observed at 35°C with 1300±60AU/ml activity followed by 30°C and 40°C with 1100±50AU/ml and 1000±50AU/ml activities and least production was recorded at both 20°C and 50°C with 200±10AU/ml activity (Fig. 5). From the above study, both the increasing and decreasing temperatures from 35°C revealed negative effects on bacteriocin synthesis. Close to this study, a probiotic L. plantarum BC-25 cultured in MRS broth revealed maximum production of bacteriocin using 35°C and pH 6.8 (Zhou et at. 2015; Zhang et al. 2020).
Figure 5: Influence of different temperature on the Bacteriocin
production using the L. plantarum MDP 5

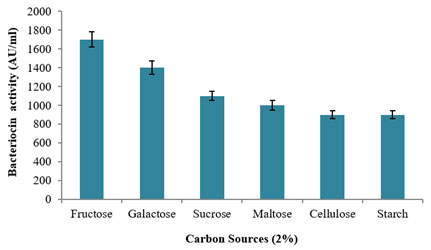
The influence on various carbon sources at 2% concentration was used for the production of bacteriocin using the potential bacterium, L. plantarum MDP 5 (Fig. 6). Among the tested carbon sources, monosaccharide substrate of fructose revealed the highest production of bacteriocin with 1700±80AU/ml activity followed by another monosaccharide substrate of galactose with 1400±70AU/ml activity. Further, a moderate bacteriocin production was observed with 1100±50AU/ml and 1000±50AU/ml while using the disaccharide sources of sucrose and maltose and a least production was evidenced using polysaccharide substrate viz., cellulose and starch in which both has 900±40AU/ml activities. However, L. acidophilus AA11 showed maximum production of bacteriocin using 0.5% lactose as its sole carbon source (Abo-Amer, 2011). In support to the present study, L. plantarum Q7 revealed appreciable production of bacteriocin using fructose in the MRS medium (Zhang et al. 2020).
Figure 6: Study of various carbon sources on the Bacteriocin
production using the L. plantarum MDP 5
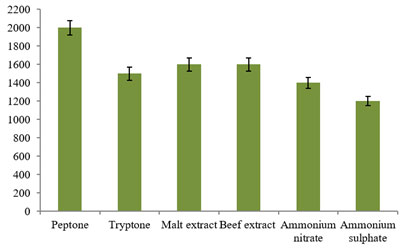
Likewise, the effect of different nitrogen sources at 1% concentration was investigated for the bacteriocin production in this study. Interestingly, organic nitrogen substrates revealed good sources for the bacteriocin production when compared to inorganic substrates used in this investigation (Fig. 7). Among them, use of peptone in the production medium evidenced maximum bacteriocin activity of 2000±80AU/ml followed by malt extract, beef extract and tryptone with 1600±80AU/ml, 1600±80AU/ml and 1500±80AU/ml activities, respectively. Further, low bacteriocin production was recorded in ammonium nitrate and ammonium sulphate with 1400±60AU/ml and 1200±50AU/ml activities, respectively.
However, L. plantarum ST13BR evidenced maximum production of bacteriocin in the combination of the meat extract and yeast extract with 1:1 ratio as its sole nitrogen source (Todorov 2004). Similarly, L. brevis C23 showed maximum bacteriocin production using meat extract as its sole nitrogen source (Sreedharan et al. 2021). From the above findings, all the four physicochemical parameters utilized in this study revealed the enhanced production of bacteriocin.
Figure 7: Study of different nitrogen sources on the Bacteriocin
production using the L. plantarum MDP 5.
CONCLUSION
The findings of the present study explored a new bacteriocin producing L. plantarum MDP 5 isolated from the milk and dairy products collected from Puducherry. The bacterium showed a growth dependent synthesis which revealed its peak production at earliest time of stationary growth phase. Further, this study evidenced enhanced bacteriocin production through the easily lab consumable physicochemical conditions which suggesting its feasibilities for commercialization opportunities. Moreover, this bacterium revealed a strong antimicrobial activity against V. cholerae MTCC 3906 which holds as baseline data for the possible future investigation regarding its antimicrobial applications against many deadly human and animal pathogens.
ACKNOWLEDGEMENTS
The study was supported by the Department of Microbiology, Faculty of Science, Annamalai University, Chidambaram, Tamil Nadu, India. Authors thank the department for providing lab facilities and supporting the research work.
Conflict of Interests: Authors declare no conflict of interest to disclose.
Data Availability Statement: The database generated and /or analysed during the current study are not publicly available due to privacy, but are available from the corresponding author on reasonable request.
REFERENCES
Abo-Amer, A.E., (2011). Optimization of bacteriocin production by Lactobacillus acidophilus AA11, a strain isolated from Egyptian cheese. Annals of Microbiology, 61(3), pp.445-452.
Ahmad, A., Banat, F. and Taher, H., (2020). Enhanced lactic acid production from food waste in dark fermentation with indigenous microbiota. Biomass Conversion and Biorefinery, pp.1-10.
Fathima, F. and Jayalakshmi, S., (2012). Characterization of urease enzyme from marine bacterium Klebsiella species. African Journal of Microbiology Research, 6(30), pp.5914-5923.
Balan, S.S., Raffi, S.M. and Jayalakshmi, S., (2013a). Probing of potential luminous bacteria in Bay of Bengal and its enzyme characterization. Journal of Microbiology and Biotechnology, 23(6), pp.811-817.
Balan, S.S. and Jayalakshmi, S., (2013b). Glycolipid biosurfactant production using low-cost medium from marine bacterium Pseudomonas aeruginosa of Mudasalodai coast. International Journal of Green Chemistry and Bioprocess, 3(3), pp.33-37.
Balan, S.S., Kumar, C.G. and Jayalakshmi, S., (2019a). Physicochemical, structural and biological evaluation of Cybersan (trigalactomargarate), a new glycolipid biosurfactant produced by a marine yeast, Cyberlindnera saturnus strain SBPN-27. Process Biochemistry, 80, pp.171-180.
Balan, S.S., Mani, P., Kumar, C.G. et al. (2019b). Structural characterization and biological evaluation of Staphylosan (dimannooleate), a new glycolipid surfactant produced by a marine Staphylococcus saprophyticus SBPS-15. Enzyme and Microbial Technology, 120, pp.1-7.
Balan, S.S., (2014). Production, characterization, evaluation of a glycolipid biosurfactant from a marine strain Bacillus cereus and development of non-toxic skin and hair care cosmetic formulations. Ph.D., Thesis, Annamalai University, Tamil Nadu, India, pp. 63-98.
Beasley, S.S. and Saris, P.E., (2004). Nisin-producing Lactococcus lactis strains isolated from human milk. Applied and Environmental Microbiology, 70(8), pp.5051-5053.
Biscola, V., Todorov, S.D., Capuano, V.S.C., et al. (2013). Isolation and characterization of a nisin-like bacteriocin produced by a Lactococcus lactis strain isolated from charqui, a Brazilian fermented, salted and dried meat product. Meat Science, 93(3), pp.607-613.
Chen, K., Liu, C., Wang, Y., et al. (2021). Predominance of indigenous non-Saccharomyces yeasts in the traditional fermentation of greengage wine and their significant contribution to the evolution of terpenes and ethyl esters. Food Research International, 143, p.110253. https://doi.org/10.1016/j.foodres.2021.110253
Cintas, L.M., Casaus, M.P., Herranz, C., et al. (2001). Bacteriocins of lactic acid bacteria. Food Science and Technology International, 7(4), pp.281-305.
Daba, G.M. and Elkhateeb, W.A., (2020). Bacteriocins of lactic acid bacteria as biotechnological tools in food and pharmaceuticals: Current applications and future prospects. Biocatalysis and Agricultural Biotechnology, p.101750. https://doi.org/ 10.1016/j.bcab.2020.101750.
Daeschel, M.A., (1989). Antimicrobial substances from lactic acid bacteria for use as food preservatives. Food Technology (Chicago), 43(1), pp.164-167.
Diop, M.B., Dauphin, R.D., Tine, E., et al. (2007). Bacteriocin producers from traditional food products. Biotechnologie, Agronomie, Société et Environnement, 11(4), pp.275-281.
Garzon, K., Ortega, C. and Tenea, G.N., (2017). Characterization of bacteriocin-producing lactic acid bacteria isolated from native fruits of Ecuadorian Amazon. Polish Journal of Microbiology, 66(4), pp.473-481.
Harley. J., (2013). Laboratory Exercises in Microbiology, 9th ed. McGraw-Hill College Division, Boston, MA.
Heredia-Castro, P.Y., Méndez-Romero, J.I., Hernández-Mendoza, A., et al. (2015). Antimicrobial activity and partial characterization of bacteriocin-like inhibitory substances produced by Lactobacillus spp. isolated from artisanal Mexican cheese. Journal of Dairy Science, 98(12), pp.8285-8293.
Jack, R.W., Tagg, J.R. and Ray, B., (1995). Bacteriocins of gram-positive bacteria. Microbiological Reviews, 59(2), pp.171-200.
Kang, J.H. and Lee, M.S., (2005). Characterization of a bacteriocin produced by Enterococcus faecium GM‐1 isolated from an infant. Journal of Applied Microbiology, 98(5), pp.1169-1176.
Klaenhammer, T.R., (1988). Bacteriocins of lactic acid bacteria. Biochimie, 70(3), pp.337-349.
Kohoutova, D., Forstlova, M., Moravkova, P., et al. (2020). Bacteriocin production by mucosal bacteria in current and previous colorectal neoplasia. BMC Cancer, 20(1), pp.1-7.
Kumar, S., Stecher, G. and Tamura, K., (2016). MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Molecular Biology and Evolution, 33(7), pp.1870-1874.
Mani, P., Sivakumar, P. and Balan, S.S., (2016). Economic production and oil recovery efficiency of a lipopeptide biosurfactant from a novel marine bacterium Bacillus simplex. Achievements in the Life Sciences, 10(1), pp.102-110.
Nebbia, S., Lamberti, C., Lo Bianco, G., et al. (2021). Antimicrobial potential of food lactic acid bacteria: bioactive peptide decrypting from caseins and bacteriocin production. Microorganisms, 9(1), p.65. https://doi.org/10.3390/microorganisms9010065
Pei, J., Jin, W., Abd El-Aty, A.M., et al. (2020). Isolation, purification, and structural identification of a new bacteriocin made by L. plantarum found in conventional kombucha. Food Control, 110, p.106923. https://doi.org/10.1016/j.foodcont.2019.106923
Rammelsberg, M. and Radler, F., (1990). Antibacterial polypeptides of Lactobacillus species. Journal of Applied Bacteriology, 69(2), pp.177-184.
Reis, J.A., Paula, A.T., Casarotti, S.N., et al. (2012). Lactic acid bacteria antimicrobial compounds: characteristics and applications. Food Engineering Reviews, 4(2), pp.124-140.
Saitou, N. and Nei, M., (1987). The neighbor-joining method: a new method for reconstructing phylogenetic trees. Molecular Biology and Evolution, 4(4), pp.406-425.
Sankar, S., Balan, S.S. and Jayalakshmi, S., (2013). Effect of biosurfactant as antibiotic role against fungal pathogens in fish Chirrinus Mirgala. International Journal of Current Trends in Research, 2(1), pp.338-344.
Sonbol, F.I., Aziz, A.A.A., El-Banna, T.E., et al. (2020). Antimicrobial activity of bacteriocins produced by Enterococcus isolates recovered from Egyptian homemade dairy products against some foodborne pathogens. International Microbiology, 23(4), pp.533-547.
Sreedharan, D. K., Abbasiliasi, S., Mohamed, M.S., et al. (2012). Fermentation strategies for improving the production of bacteriocin‐like inhibitory substances by Lactobacillus brevis C23 with nutrient supplementation, pH and temperature variations. Journal of Food Processing and Preservation, p.e15914. https://doi.org/10.1111/jfpp.15914
Todorov, S.D. and Dicks, L.M.T., (2006). Screening for bacteriocin-producing lactic acid bacteria from boza, a traditional cereal beverage from Bulgaria: Comparison of the bacteriocins. Process Biochemistry, 41(1), pp.11-19.
Todorov, S.D., van Reenen, C.A. and Dicks, L.M.T., (2004). Optimization of bacteriocin production by Lactobacillus plantarum ST13BR, a strain isolated from barley beer. The Journal of General and Applied Microbiology, 50(3), pp.149-157.
Zhang, J., Han, X., Zhang, L., et al. (2020). Effects of fructose and overexpression of shock-related gene grol on plantaricin Q7 production. Probiotics and Antimicrobial Proteins, 12(1), pp.32-38.
Zhou, K., Zeng, Y.T., Han, X.F. et al. (2015). Modelling growth and bacteriocin production by Lactobacillus plantarum BC-25 in response to temperature and pH in batch fermentation. Applied Biochemistry and Biotechnology, 176(6), pp.1627-1637.


