1Department of Biotechnology, National College, (Autonomous), Tiruchirappalli, Tamil Nadu, India
2Department of Biosciences, Integral University Lucknow, Uttar Pradesh India
Corresponding author Email: mohamedjaabir@nct.ac.in
Article Publishing History
Received: 02/01/2019
Accepted After Revision: 07/03/2019
The increasing cases of resistance, low effi cacy and high toxicity of antibiotics and anticancer agents have led to the discovery of nanoparticles as potent antimicrobial and anticancer agents to combat the threat of Multi Drug Resistance (MDR) and to minimize the side effects of drugs. The synthesized silver nanoparticles (SAgNPs) (synthesized using MDR Staphylococcus aureus) were characterized to confi rm synthesis, shape, size, hydrodynamic diameter, colloidal stability and surface functionalization, by UV-visible spectroscopy, TEM, DLS, zeta potential test and FTIR respectively. SAgNPs were found to have spherical shape with a size of 15 nm. Hydrodynamic diameter of nanoparticles was found to be 88.65 nm and the value of zeta potential was recorded to be -24.03 mV. The antimicrobial and anticancer potential of SAgNPs was performed against normal & MDR strains of S. aureus and human colon cancer cell line (HCT-116), respectively.. The MICs of SAgNPs against normal and MDR S. aureus strains were found to be 0.025 μg/ml & 0.053 μg/ ml, respectively. Similarly, IC50 against HCT-116 cell line was found to be 0.069 μg/ml by MTT Assay. DAPI analysis confi rmed the interaction of SAgNPs with DNA in order to initiate apoptosis for killing cancer cells.
Anticancer; Dapi; Dls; Mdr; Mtt; Sagnps
Haseeb M, Khan M. S, Baker A, Khan I, Wahid I, Jaabir M. S. M. Anticancer and Antibacterial Potential of MDR Staphylococcus Aureus Mediated Synthesized Silver Nanoparticles. Biosc.Biotech.Res.Comm. 2019;12(1).
Haseeb M, Khan M. S, Baker A, Khan I, Wahid I, Jaabir M. S. M. Anticancer and Antibacterial Potential of MDR Staphylococcus Aureus Mediated Synthesized Silver Nanoparticles. Biosc.Biotech.Res.Comm. 2019;12(1). Available from: https://bit.ly/2M4QoFl
Copyright © Haseeb et al., This is an open access article distributed under the terms of the Creative Commons Attribution License (CC-BY) https://creativecommns.org/licenses/by/4.0/, which permits unrestricted use distribution and reproduction in any medium, provide the original author and source are credited.
Introduction
Toxicity, high cost of the drugs used in the treatment of cancer and microbial diseases, rise in the cases of antibiotic resistance due to frequent use of antibiotics have forced researchers to look for alternatives in medicine and therapeutics. Side effects of chemotherapeutic drugs due to poor specifi city towards targets and high dose requirements have also made it necessary for nano-medicine to address the issue and look for other efficient and cost effective molecules bearing high degree of effi cacy for the target cells (Riehemann et al., 2009).
Inorganic nanoparticles are being widely used throughout the world due to their incredible yet, potential applications. Among the metal NPs, Silver nanoparticles (AgNPs) in particular have attracted considerable attention in various fields and have been used as therapeutic agent (Shrivastava et al., 2009), as catalyst (Christopher et al., 2011) antimicrobial and anti-infl ammatory agents (El-Chaghaby, and Ahmad, 2011; Veerasamy et al., 2011), as a strong cytotoxic agent against cancer cell lines (Jacob et al., 2012) . Silver and silver ions have been used since a long time as strong antimicrobial and anti-infl ammatory agents. Nano-silver or silver nanoparticles (AgNPs) have shown promising results against the antibiotic resistant bacteria (Yah and Simate, 2015). AgNPs have also been investigated for their antimicrobial potential against Multi Drug Resistant MDR strains of several pathogenic microbes (Alshaye et al., 2017). The role of AgNPs in diagnostic and probing of cancer has also been reported earlier in a research study (Huang et al., 2017).
In literature, several chemical and physical methods have been used for the synthesis of AgNPs, but these methods utilize large amount of toxic chemicals and extreme conditions like high temperature and are not economical. The green synthesis approaches are a solution to this problem and are more ecofriendly, simple, reliable, reproducible and non-toxic. Recently, researchers have exploited microorganism as a production tool for the biosynthesis of inorganic metallic nanoparticles, such as silver, cadmium, gold and sulfi de (Hassaan et al., 2018; Elsalam et al., 2018).
The bacterial synthesis seems to be advantageous because of minimal utilization of toxic chemicals and sustainability of large scale production. Many reports have shown that numerous bacterial strains of Staphylococcus aureus, Bacillus cereus, Bacillus subtilis, Escherichia coli, Bacillus licheniformis, Enterobacter cloacae, Klebsiella pneumonia, Lactobacillus acidophilus, and Pseudomonas aeroginosa, are capable of synthesizing nanoparticles (NPs) (Ghiua˘ et al., 2018; Muthulakshmi et al. 2018; Fatemi et al., 2018).
Biosynthetic methods can be categorized into two groups, on the basis of location of synthesis i.e. intracellular and extracellular synthesis. The extracellular synthesis of nanoparticles is simple and economical because of the simplicity in its procedure for not only synthesis but also of recovery and purifi cation as well, under large scale production (Singh et al., 2018). All these reasons make the bacteria a potential source for the extracellular synthesis of AgNPs avoiding the use of toxic and hazardous chemicals.
Novel technologies are being developed to overcome the challenges imposed by bacteria and cancer cells, the resistance associated with the regular use of antibiotics through MDR phenomenon and to minimize the side effects of the drugs used in chemotherapy (Mohammed et al., 2018).
The current study aims to address the issues of MDR resistance of bacteria S. aureus an uses AgNPs as a tool against cancer cells. Here, AgNPs were synthesized extracellularly from bacteria MDR S. aureus, characterized by UV-vis spectroscopy, Transmission Electron Microscopy (TEM), Zeta potential value, Dynamic Light Scattering (DLS), Fourier Transformed Infra Red (FTIR), and subsequently tested against normal and MDR S. aureus as well as against cancer cell lines HCT-116.
Materials and Methods
All the chemicals and media were purchased from Sigma Aldrich (St. Louis, USA) and HiMedia, India. The multidrug resistant (MDR) and normal strain of Staphylococcus aureus (NCIM 2079) were obtained from NCIM, Pune, India.
Synthesis of AgNPs
The MDR strain of S. aureus (NCIM 2079) was used for the synthesis of AgNPs. Bacterial strain was maintained on nutrient agar at 37o C. The bacterial cultures (OD600- 0.60) were transferred to Erlenmeyer fl asks containing 250 ml nutrient broth and incubated at 37oC on a rotary shaker (180 rpm) for 6-8 hrs. The bacterial cells were collected by centrifugation (6000 g, 10 min at 10°C), washed extensively with sterile distilled water under aseptic conditions and used for further studies. Two different reactions were carried out by incubating S. aureus (NCIM 2079) cells (wet weight-5gm) in 100ml sterile distilled water containing 1mM AgNO3 in two different 500ml Erlenmeyer fl asks for 6-8 hrs at rotatory shaker (180 rpm) at 37°C. The monitoring of the bacteria mediated reduction of silver ions was accomplished by the visual color change and UV–visible spectrum analysis for the reaction mixture.
At the end of synthesis, the unbound proteins were removed by Precipitation with 100% (v/v) volumes of absolute ethanol and the AgNPs were collected for further characterization.
Characterization of AgNPs
The AgNPs synthesized using S.aureus (NCIM 2079) were characterized by UV-Vis Spectroscopy, Dynamic Light Scattering, Zeta potential analysis, TEM and FTIR for the confi rmation of synthesis, size, shape, stability, surface functionalization and identifi cation.UV-Vis spectrophotometer measurements were performed on a Shimadzu dual-beam spectrophotometer (model UV-1601 PC) operated at a resolution of 1nm.
Transmission Electron Microscopy (TEM) measurements were performed to analyze the size and morphology. Samples were prepared by drying a drop of AgNPs solution on carbon coated TEM copper grids followed by measurements on (TEM) FEI Company, TecnaiTM G2 Spirit BioTWIN operated at an accelerating voltage of 80kV. The study was done Indian Institute of Toxicological Research (IITR), Lucknow, India. Dynamic Light Scattering (DLS) was used to analyze the mean particle size (MPS) of biosynthesized AgNPs using dynamic light scattering particle size analyzer (Zetasizer Nano- ZS, Model ZEN3600, Malvern Instrument Ltd, Malvern, UK). The samples were taken in a DTS0112-low volume disposable sizing cuvette of 1.5 ml capacity. The sample powder was dissolved to a concentration of 0.5% (w/v) in deionised water and sonicated for 1 min before analysis. After fi ltering through a syringe fi lter of 0.4μm pore size, solution was centrifuged at 5000 rpm for 30min and measured for its particle size. Mean particle size was the average of triplicate measurements for a single sample. Zeta potential of AgNPs was determined to measure the charge, as the metal nanoparticles are normally charged or carry charge of capping agents. This analysis was done in Zetasizer Nano-ZS, (Malvern Instrument Ltd. and Malvern, UK).Fourier-transform infrared spectroscopy analysis (FTIR) was used to identify capped biomolecules over the surface of as synthesized AgNPs was done by FT-IR (Perkin Elmer Spectrum, Jasco- 6100). For the analysis, silver nanoparticles were dried, grounded with KBr pellets and analyzed in the wavelength range of 4000 to 400 cm–1.
The antibacterial activity of AgNPs was determined against MDR strain of Staphylococcus aureus (NICM 2079), and Normal Strains of Staphylococcus aureus (NICM 2079). The bacterial cells (OD600-0.60) were allowed to grow in nutrient broth for 24 hours and kept in a shaker incubator at 180 rpm for 24 hrs at 37C. Bacterial lawns were prepared using 100 ml of cultured broth. Agar well diffusion method (Khan et al., 2011) was used to determine preliminary antibacterial activity of biosynthesized AgNPs. Plates were prepared by pouring around 25 ml of sterile Mueller Hinton Agar (MHA) media in sterilized petridishes. Different strains of bacterial culture chosen for the study was swabbed using sterilized cotton swab. Sterilized gel puncture was used to make wells of 5 mm diameter. Two wells were made in each plate, with each one containing distilled water (for control) and the other was loaded with synthesized silver nanoparticles. The plates were left for 24 hr incubation at 37C. The inhibition was examined by identifying the clear zone around the well.
The minimum inhibitory concentration (MIC) of the synthesized AgNPs was determined by double diffusion method (Sarker et al., 2007). The MIC50 was determined by broth dilution, performed on a 96 fl at-bottom well plate. The medium used in the plates were prepared at double the fi nal strength to allow for a 50% dilution once the inoculum is added. This approach allowed the inoculum to be prepared in distilled water, which permitted the absorbance to be determined using a spectrophotometer without interference from colored media. All the bacterial strains were allowed to grow till midlogrithmic phase. Culture was subsequently harvested by centrifugation, washed with 1mM sodium Phosphate buffer (SPB) at pH 7.4, and diluted to 2 × 105 colony forming units (CFU)/ml in Saline phosphate buffer. About 90 μL of Mueller-Hinton broth was used in 96 well- microtitre plates to serially dilute the silver nanoparticles in desired concentrations. Bacterial suspension of 95×104 CFU/ well was used as inoculum. Inoculated Microtitre wells were incubated overnight at 37o C. The MIC50 was determined by taking lowest concentration of AgNPs at which the bacterial growth was inhibited. The numbers of colonies were determined by agar plate count method as discussed by Morones et al., (2005).The minimum bactericidal concentration (MBC) is the lowest concentration of an antibacterial agent required to kill a particular bacterium. It can be determined from broth dilution minimum inhibitory concentration (MIC) tests by sub culturing to agar plates that do not contain the test agent.
Cell viability and IC50 value determination of AgNPs against cancer line HCT-116
The biosynthesized silver nanoparticles were tested against the Human colon cancer cell line HCT-116. The anticancer activity of synthesized silver NPs was determined by MTT (3-[4, 5-dimethylthiazol-2-yl]-2, 5-diphenyltetrazolium bromide) assay and nuclear degradation was checked by DAPI (4’, 6-diamino-2-phenylindiole) toxicology assays. For this, the cells were treated with varying concentration of AgNPs and the effect was analyzed by MTT assay and DAPI.
To determine the effect of Ag nanoparticles on cancer cells, MTT assay was performed. For this the HCT 116 cells were plated in 96 well plate with each well seeded with 1 X 104 cells. The plate was incubated for 24 hours in CO2 incubator. After incubation, cells were treated with Ag NP of various concentrations in triplicate with untreated cells as the positive control and incubated for 48 hours. After this, media was removed and wells were loaded with 50μl of 3-(4, 5-dimethylthiazol-2-yl)-2, 5-diphenyltetrazolium bromide (MTT) dye (5mg/ml in PBS) and fresh culture media. Then the plate was incubated for 4 hours. Formosan crystals formed during the process were dissolved by adding 100μl of DMSO (Dimethyl sulfoxide) and incubated for one hour. The reduced MTT was quantifi ed by measuring optical density at 570 nm in ELISA reader.
The Percentage inhibition of the cells was calculated using the formula
X = 100 − (A test – A blank) / (A control – A blank) × 100
Where, X- Percentage inhibition, A test − absorbance of the test sample, A blank − absorbance of blank and A control − absorbance of the control sample. The IC50 value was calculated from the data obtained.
The Apoptotic effect of Ag NP on the cell line was determined by nuclear fl uorescent staining by DAPI. For this the cells were seeded in 96 well plate and treated with the Ag NP as mentioned above. Then media was removed and cells were washed with PBS and fi xed with 4% para formaldehyde for 10 minutes. Then the staining with DAPI was performed along with permeabilizing buffer to fi x the stain into the cells. After staining, the cell imaging was obtained under fl uorescence microscope. The cells showing fragmented and condensed nuclei were considered as the apoptotic cells.
Results and Discussion
Nanoparticles synthesis was observed as the color of reaction mixture changed from light yellow to dark brown, a characteristic color change associated with formation of silver nanoparticles. UV-Vis, spectroscopy is simple and sensitive techniques for characterization of colloidal suspension and requires very short period of time for measurement (Tomaszewska et al., 2013). In UV-Vis, spectroscopy, the measurement of intensity of light passing through the sample is done, as the optical properties of metal nanoparticles are due to collective oscillation of conduction electrons, excited by electromagnetic radiation (Singh et al., 2018).
The UV-Vis spectrum in our results shows the maximum absorbance peak around 430 nm, thereby confi rming the synthesis of AgNPs in the sample.
Transmission Electron Microscopy (TEM) was used to visualize the shape and to determine the size distribution of synthesized AgNPs (fi g.2.). TEM images were obtained using JEOL 3010, operating at 200 kV accelerating voltage. The TEM analysis was performed at Indian Institute of Toxicological Research (IITR), Lucknow, India. The high resolution images for topographical studies revealed the structure and morphology of synthesized silver nanoparticles (AgNPs) and average size was found to be 15 nm.
Gomaa and Zakaria (2017) have reported similarly in the synthesis of spherical shaped AgNPs with an average size of 17 nm using Staphylococcus aureus and Escherichia coli.
Dynamic light scattering (DLS) based on the laser diffraction method with multiple scattering technique was chosen to determine the size of AgNPs. DLS is a method which is dependent upon the interaction of light with particles in suspension. It is typically used to measure the particles size distribution in the range of 2 nm to 500 nm (Tomaszewska et al., 2013). In DLS measurement the scattered light passing through the colloidal suspension relies on Rayleigh scattering from the particles in suspension (Fissan et al., 2014). Next, the hydrodynamic size of the particles is determined by analyzing the modulation of intensity of scattered light as function of time (Dieckmann et al., 2009). DLS revealed the Z-average
value to be 88.65 nm. Similar results were obtained in another research in which average diameter silver NPs was found to be 87.46 nm in the aqueous colloidal suspension of AgNPs. (Singh et al., 2018).
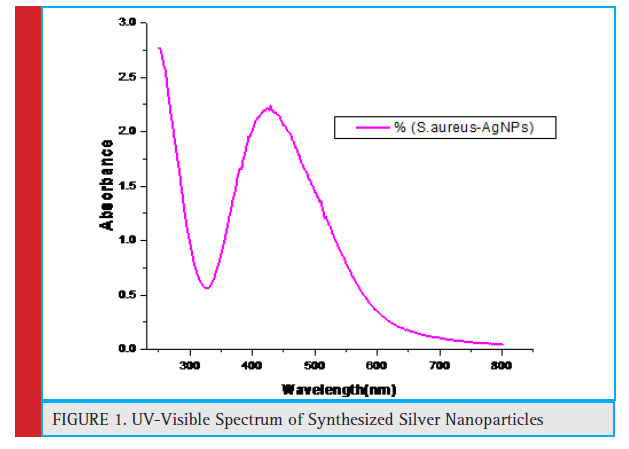 |
Figure 1: UV-Visible Spectrum of Synthesized Silver Nanoparticles |
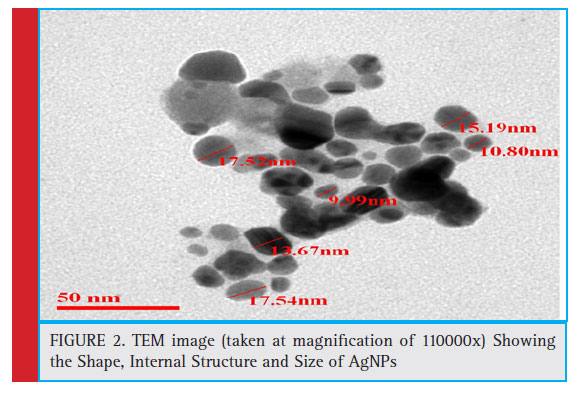 |
Figure 2: TEM image (taken at magnifi cation of 110000x) Showing the Shape, Internal Structure and Size of AgNPs |
The determination of Zeta potential is considered as an effective simplest, and most straight forward method to predict the stability and understand surface properties of the nanoparticles. Zeta potential is a measure of the colloidal stability of the nanoparticles in a solution (Saeb et al., 2014). The shielding or exposure of charged groups or concentration, distribution, ionization and adsorption of nano-particles can also be estimated from zeta potential (Guilatt et al., 2004).
Information with reference to the concentration, distribution, exposure or shielding of charged moieties; ionization and adsorption could be inferred from the analysis of zeta potential. In the present study, the zeta potential of the synthesized AgNPs was found to be -24.3 mV, which confi rmed that the AgNPs were highly stable in colloidal suspension. Similar observation were also recorded in which zeta potential value was measured to be -25.5 mV (Singh et al., 2018).
FTIR of the synthesized AgNPs is performed to spot the position of various functional groups of the capping agent and their vibration patterns. The FTIR results of the sample states that the position of Amide I(C=O) bond is 1637 cm-1 with transmittance of 10.75%, O-H(Stretch) is 3583.13 cm-1 with transmittance of 9.19%, C-O(Stretch) is 1082.41 cm-1 with transmittance of 16.19% and Alkyne (Stretch) bond 2097.88 cm-1 with transmittance of 33.17%.
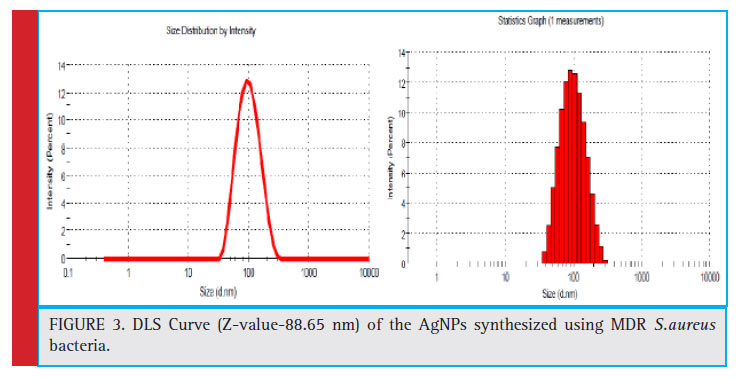 |
Figure 3: DLS Curve (Z-value-88.65 nm) of the AgNPs synthesized using MDR S.aureus bacteria. |
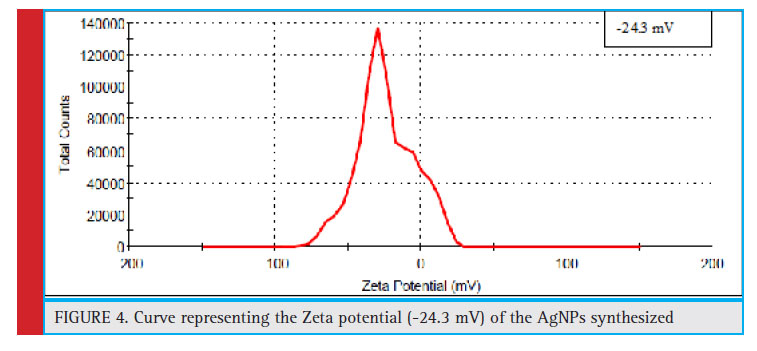 |
Figure 4: Curve representing the Zeta potential (-24.3 mV) of the AgNPs synthesized |
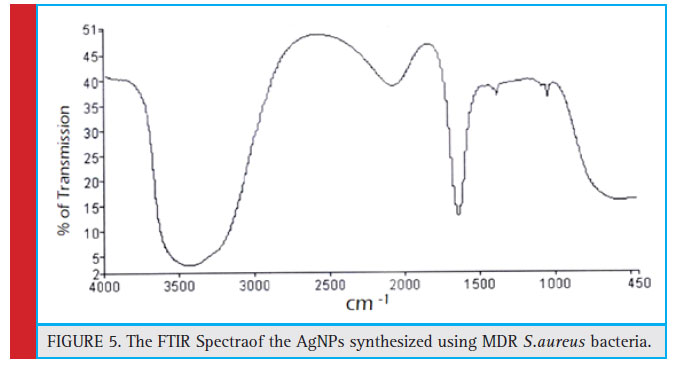 |
Figure 5: The FTIR Spectraof the AgNPs synthesized using MDR S.aureus bacteria |
Antibacterial activity of AgNPs against normal and MDR strains
The AgNPs synthesized using MDR S.aureus was tested for its antibacterial potential, against normal strains of S.aureus. The same was tested against strains of MDR S.aureus as well. The antimicrobial potential was tested by agar well diffusion method, MIC and MBC.
In general MIC is defi ned as the minimum amount of drug required to inhibit the growth of bacteria to 70%- 80%. MIC50 is defi ned as the minimum amount of drug required to inhibit the growth of bacteria to 50%. The clear zone of inhibition around the inoculated area confi rmed the antibacterial potential of as the synthesized AgNPs. MIC and MBC concentrations were calculated under varying concentrations of the AgNPs synthesized using MDR S.aureus. The Zone of inhibition of AgNPs against normal strain and MDR strains of S.aureus were found to be 2.8 cm and 2.5 cm respectively (figure no. 6a and 6b). The Minimum inhibitory concentration of AgNPs against normal strain and MDR strains of S.aureus were found to be 0.025 μg/ml and 0.053 μg/ ml for respectively. The lowest concentration with no visible growth indicating 99.5% killing of the inoculum
was found to be double the value of MIC, (see figure 8 a and b).
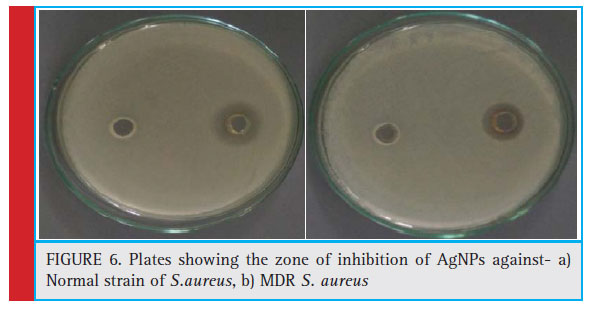 |
Figure 6: Plates showing the zone of inhibition of AgNPs against- a) Normal strain of S.aureus, b) MDR S. aureus |
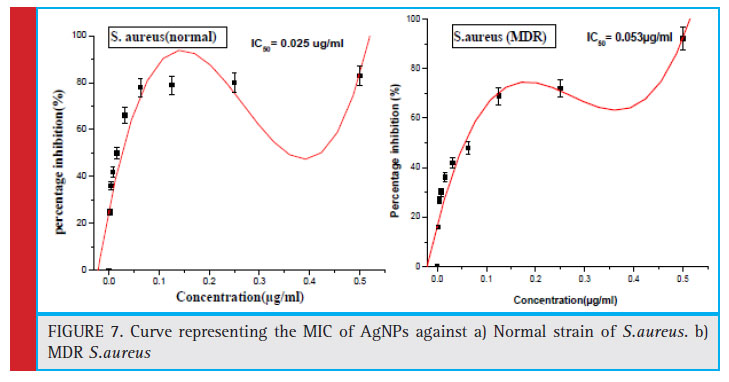 |
Figure 7: Curve representing the MIC of AgNPs against a) Normal strain of S.aureus. b) MDR S.aureus |
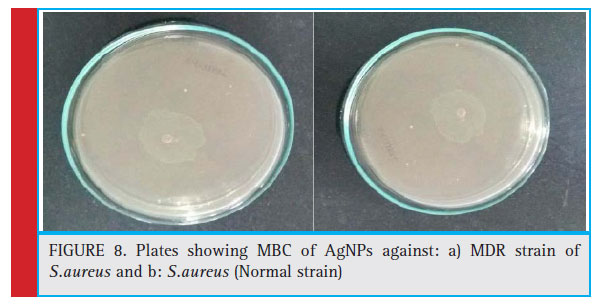 |
Figure 8: Plates showing MBC of AgNPs against: a) MDR strain of S.aureus and b: S.aureus (Normal strain) |
Anticancer activity against HCT-116
To evaluate the sensitivity of colon cancer cells to the AgNPs, human colon cancer cell line HCT-116 was treated with different doses (0.01–10 μg/ml) of AgNPs synthesized using MDR S.aureus for 24 and 48 h followed by MTT assay (Fig. 9 a and 9b). MTT is a dye which is reduced to formazon crystals during metabolism, cellular mechanism of MTT reduction into formazan is understood, involving reaction with NADH or similar reducing molecules that transfer electrons to MTT Viable cells with active metabolism convert MTT into a purple colored formazan product with an absorbance maximum near 570 nm. When cells die, they lose the ability to convert MTT into formazan, thus color formation serves as a useful and convenient marker of only the viable cells. So, the intensity of the colored product is measured which is directly proportional to the number of viable cells in the culture (Präbst et al., 2017). The MTT analysis was done to assess the activity of mitochondrial succinate dehydrogenase (Singh et al., 2018). Here, the decrease in the MTT reduction could be due to decrease in cell viability which in turn can be attributed to cytotoxic activity of silver nanoparticles. Our results shows that AgNPs at a concentration of 0.069 μg/ml reduced growth of human colon cancer cells by 50%, after 24 h of treatment. Similar pattern of results with more pronounced cytotoxic effect was observed after 48 h of treatment thus proving AgNPs to be more anti-proliferative and cytotoxic for colon cancer cells. Similar fi ndings have been reported in earlier studies, (Singh et al., 2018; Patra et al., 2018).
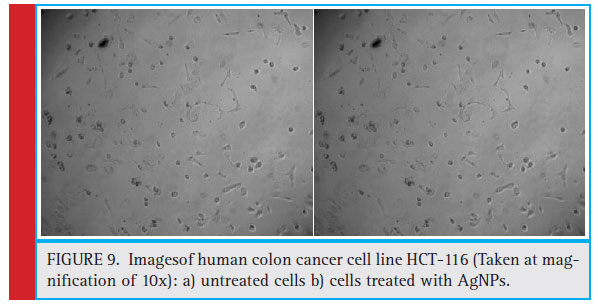 |
Figure 9: Imagesof human colon cancer cell line HCT-116 (Taken at magnifi cation of 10x): a) untreated cells b) cells treated with AgNPs. |
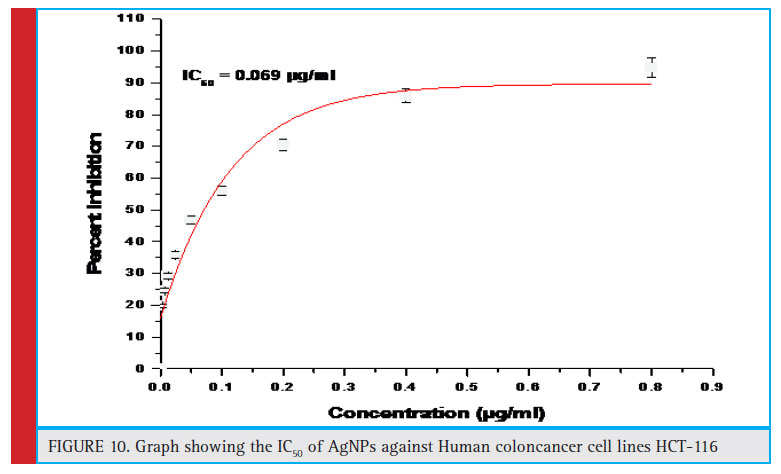 |
Figure 10: Graph showing the IC50 of AgNPs against Human coloncancer cell lines HCT-116 |
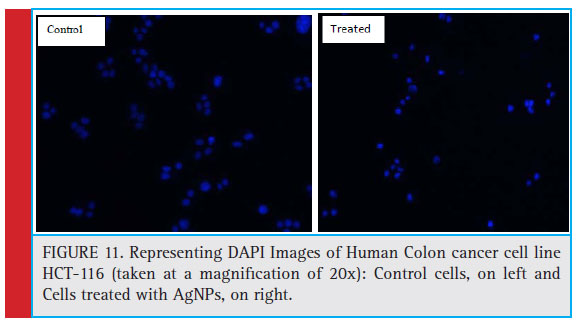 |
Figure 11: Representing DAPI Images of Human Colon cancer cell line HCT-116 (taken at a magnifi cation of 20x): Control cells, on left and Cells treated with AgNPs, on right. |
AgNPs induced nuclear condensation in HCT-116 cells: Apoptosis is characterized by changes in cellular morphology such nuclear fragmentation, degradation of DNA, condensation of chromatin, apoptotic body formation and membrane blabbing (Sheikh et al., 2018). In order to confi rm apoptosis in AgNPs treated colon cancer cells, DAPI nuclear staining was performed. DAPI (4’, 6-diamidino-2-phenylindole) is a fl uorescent stain which preferably binds to AT rich regions of DNA.
The fl uorescence intensity of DAPI increases to approximately 20 times when DAPI is bound to DNA. As the ell permeability is compromised due to apoptosis; DAPI enters easily inside the cell where its binding to DNA produces a strong blue color (Baharara et al., 2018).
After treatment with AgNPs for 24 h, signifi cant nuclear changes in colon cancer cells were observed via DAPI staining. As apparent from photomicrographs, AgNPs induced nuclear condensation and fragmentation in colon cancer cells in a dose dependent manner, whereas the control cells exhibited normal cell morphology. The results were evident that AgNPs induced apoptosis in colon cancer cells in a time and dose-dependent manner. Our results are in agreement with several other researches published earlier (Gurunathan et al., 2018; Baharara et al., 2018; Kishore et al., 2018).
Conclusion
In this study, MDR strain of S. aureus was used to synthesize silver NPs, with the goal to inhibit the MDR strains of S.aureus as well as normal strain of S.aureus to assess its anticancer potential.The synthesized NPs were found to be spherical in shape and approximately 15 nm in size. The NPs were found to form a stable suspension with a zeta potential value of -24.03 m. These particles were found to be capped by extracellular bacterial proteins (confi rmed by FTIR). The minimum inhibitory concentrations were found to be 0.025 μg/ml for S.aureus (normal) and 0.053 μg/ml for S.aureus (MDR), respectively. These nanoparticles were also found to be substantially effective against human colon cancer cell lines HCT-116, with an IC50 value of 0.069 μg/ml. Further optimization is needed in order to assess the anticancer properties of as synthesized AgNPs.
Acknowledgement
Authors acknowledge the support of National College (Autonomous) for providing all the necessary facilities to carry out this research work.
References
Alshaye, N.A., Elobeid, M.M., Alkhalifah, D.H. and Mohammed, A.E., (2017). Characterization of Biogenic Silver Nanoparticles by Salvadora persica Leaves Extract and its Application Against Some MDR Pathogens E. coli and S. aureus. Research Journal of Microbiology, 12, pp.74-81.
Baharara, J., Ramezani, T., Hosseini, N. and Mousavi, M., (2018). Silver Nanoparticles Synthesized Coating with Zatariamultifl ora Leaves Extract Induced Apoptosis in HeLa Cells Through p53 Activation. Iranian journal of pharmaceutical research: IJPR, 17(2), p.627.
Christopher, P., Xin, H. and Linic, S., (2011). Visible-lightenhanced catalytic oxidation reactions on plasmonic silver nanostructures. Nature chemistry, 3(6), p.467.
Dieckmann, Y., Cö lfen, H., Hofmann, H. and Petri-Fink, A., (2009). Particle size distribution measurements of manganese- doped ZnS nanoparticles. Analytical chemistry, 81(10), pp.3889-3895.
Elsalam, S.S.A., Taha, R.H., Tawfeik, A.M., El-Monem, M.O.A. and Mahmoud, H.A., (2018). Antimicrobial Activity of Bio and Chemical Synthesized Cadmium Sulfi de Nanoparticles. Egyptian Journal of Hospital Medicine, 70(9).
El-Chaghaby, G.A. and Ahmad, A.F., (2011). Biosynthesis of silver nanoparticles using Pistacia lentiscus leaves extract and investigation of their antimicrobial effect. Oriental Journal of Chemistry, 27(3), p.929.
Fatemi, M., Mollania, N., Momeni-Moghaddam, M. and Sadeghifar, F., (2018). Extracellular biosynthesis of magnetic iron oxide nanoparticles by Bacillus cereus strain HMH1: Characterization and in vitro cytotoxicity analysis on MCF-7 and 3T3 cell lines. Journal of biotechnology, 270, pp.1-11.
Fissan, H., Ristig, S., Kaminski, H., Asbach, C. and Epple, M., (2014). Comparison of different characterization methods for nanoparticle dispersions before and after aerosolization. Analytical Methods, 6(18), pp.7324-7334.
Ghiua˘ , I., Cristea, D., Croitoru, C., Kost, J., Wenkert, R., Vyrides, I., Anayiotos, A. and Munteanu, D., (2018). Characterization and antimicrobial activity of silver nanoparticles, biosynthesized using Bacillus species. Applied Surface Science, 438, pp.66-73.
Gomaa, E.Z., (2017). Silver nanoparticles as an antimicrobial agent: A case study on Staphylococcus aureus and Escherichia coli as models for Gram-positive and Gram-negative bacteria. The Journal of general and applied microbiology, 63(1), pp.36-43.
Gurunathan, S., Qasim, M., Park, C., Yoo, H., Kim, J.H. and Hong, K., (2018). Cytotoxic Potential and Molecular Pathway Analysis of Silver Nanoparticles in Human Colon Cancer Cells HCT116. International journal of molecular sciences, 19(8), p.2269.
Hassaan, M.A. and Hosny, S., (2018). Green Synthesis of Ag and Au Nanoparticles from Micro and Macro Algae-Review. International
Journal of Atmospheric and Oceanic Sciences, 2(1), p.10.
Huang, Y., Fan, C.Q., Dong, H., Wang, S.M., Yang, X.C. and Yang, S.M., (2017). Current applications and future prospects of nanomaterials in tumor therapy. International journal of nanomedicine, 12, p.1815.
Jacob, S.J.P., Finub, J.S. and Narayanan, A., (2012). Synthesis of silver nanoparticles using Piper longum leaf extracts and its cytotoxic activity against Hep-2 cell line. Colloids and Surfaces B: Biointerfaces, 91, pp.212-214.
Kishore, M., Abdulqader, A.T., Ahmad, H.S. and Hanumantharao, Y., (2018). Anticancer and antibacterial potential of green silver nanoparticles synthesized from Maytenus senegalensis (L.) leaf extract and their characterization. Drug Invention Today, 10(4).
Mohammed, A.E., Al-Qahtani, A., Al-Mutairi, A., Al-Shamri, B. and Aabed, K.F., (2018). Antibacterial and Cytotoxic Potential of Biosynthesized Silver Nanoparticles by Some Plant Extracts. Nanomaterials (Basel, Switzerland), 8(6).
Morones, J.R., Elechiguerra, J.L., Camacho, A., Holt, K., Kouri, J.B., Ramírez, J.T. and Yacaman, M.J., (2005). The bactericidal effect of silver nanoparticles. Nanotechnology, 16(10), p. 2346.
Muthulakshmi, K., Uma, C., Sivagurunathan, P., Yoganathan, K. and Satheeshkumar, S., (2018). Extracellular, biosynthesis of silver nanoparticles using Enterobacter cloacae (mk163462) and their antibacterial activity against certain multidrug resistant pathogens. Journal of Pharmacognosy and Phytochemistry, 7(6), pp.741-747.
Patra, N., Kar, D., Pal, A. and Behera, A., 2018. Antibacterial, anticancer, anti-diabetic and catalytic activity of bio-conjugated metal nanoparticles. Advances in Natural Sciences: Nanoscience and Nanotechnology, 9(3), p.035001.
Präbst, K., Engelhardt, H., Ringgeler, S. and Hübner, H., (2017). Basic colorimetric proliferation assays: MTT, WST, and resazurin.Cell Viability Assays: Methods and Protocols, pp.1-17.
Rabinovich-Guilatt, L., Couvreur, P., Lambert, G., Goldstein, D., Benita, S. and Dubernet, C., (2004). Extensive surface studies help to analyse zeta potential data: the case of cationic emulsions. Chemistry and Physics of Lipids, 131(1), pp.1-13.
Riehemann, K., Schneider, S.W., Luger, T.A., Godin, B., Ferrari, M. and Fuchs, H., (2009). Nanomedicine—challenge and perspectives. Angewandte Chemie International Edition, 48(5), pp.872-897.
Saeb, A., Alshammari, A.S., Al-Brahim, H. and Al-Rubeaan, K.A., (2014). Production of silver nanoparticles with strong and stable antimicrobial activity against highly pathogenic and multidrug resistant bacteria. The Scientifi c World Journal, 2014.
Sarker, S.D., Nahar, L. and Kumarasamy, Y., (2007). Microtitre plate-based antibacterial assay incorporating resazurin as an indicator of cell growth, and its application in the in vitro antibacterial screening of phytochemicals. Methods, 42(4), pp.321-324.
Sheikh, E., Bhatt, M.B. and Tripathi, M., (2018). Bio-based synthesised and characterized monodispersed Curcuma longa silver
nanoparticles induce targeted anticancer activity in breast cancer cells. Pharmacognosy Magazine, 14(57), p.340.
Shrivastava, S., Bera, T., Singh, S.K., Singh, G., Ramachandrarao, P. and Dash, D., (2009). Characterization of antiplatelet properties of silver nanoparticles. ACS nano, 3(6), pp.1357- 1364.
Singh, H., Du, J., Singh, P. and Yi, T.H., (2018). Ecofriendly synthesis of silver and gold nanoparticles by Euphrasia offi cinalis leaf extract and its biomedical applications. Artifi cial cells, nanomedicine, and biotechnology, 46(6), pp.1163-1170.
Singh, H., Du, J., Singh, P. and Yi, T.H., (2018). Extracellular synthesis of silver nanoparticles by Pseudomonas sp. THG-LS1. 4 and their antimicrobial application. Journal of pharmaceutical analysis, 8(4), pp.258-264.
Singh, H., Du, J., Singh, P. and Yi, T.H., (2018). Role of green silver nanoparticles synthesized from Symphytum offi cinale leaf extract in protection against UVB-induced photoaging. Journal of Nanostructure in Chemistry, 8(3), pp.359-368.
Sudheer Khan, S., Bharath Kumar, E., Mukherjee, A. and Chandrasekaran, N., (2011). Bacterial tolerance to silver nanoparticles (SNPs): Aeromonas punctata isolated from sewage environment. Journal of basic microbiology, 51(2), pp.183-190.
Tomaszewska, E., Soliwoda, K., Kadziola, K., Tkacz-Szczesna, B., Celichowski, G., Cichomski, M., Szmaja, W. and Grobelny, J., (2013). Detection limits of DLS and UV-Vis spectroscopy in characterization of polydisperse nanoparticles colloids. Journal of Nanomaterials 13, p.60.
Veerasamy, R., Xin, T.Z., Gunasagaran, S., Xiang, T.F.W., Yang, E.F.C., Jeyakumar, N. and Dhanaraj, S.A., 2011. Biosynthesis of silver nanoparticles using mangosteen leaf extract and evaluation of their antimicrobial activities. Journal of Saudi Chemical Society, 15(2), pp.113-120.
Yah, C.S. and Simate, G.S., 2015. Nanoparticles as potential new generation broad spectrum antimicrobial agents. DARU Journal of Pharmaceutical Sciences, 23(1), p.43.
Zweifel, U.L. and Hagstrom, A., 1995. Total counts of marine bacteria include a large fraction of non-nucleoid-containing bacteria (ghosts). Applied and Environmental Microbiology, 61(6), pp.2180-2185.


