Department of Biotechnology, Mehsana Urban Institute of Sciences, Ganpat University
Corresponding author email: priti.patel@ganpatuniversity.ac.in
Article Publishing History
Received: 05/03/2021
Accepted After Revision: 10/06/2021
Intensity of colour change from yellow to brown was measured with Shimadzu UV-Vis 1601 spectrophotometer. Size distribution and Zeta potential of silver nanoparticles were known with Malvern, and TEM analysis was done to have a detailed account of prepared silver nanoparticles’ characteristics. The diffusion method was used for analyses of efficiencies shown by the prepared silver nanoparticles and the antibiotics against various human pathogens. In result, the zones of inhibition were formed with strain synthesized silver nanoparticles as 22 mm, 24 mm, 25 mm and 24 mm for Salmonella, Co-agulase negative Staphylococcus, Klebsiella, and Enterococcus respectively. In conclusion, silver nanoparticles synthesized by Streptomyces spp. AK3 have been found a better alternative to antibiotics in terms of their efficiencies against human pathogens and side effects.
Silver Nanoparticles, Streptomyces spp. AK3, TEM Analysis, Zeta Potential Analysis, Zones of Inhibition
Kulshrestha A. K, Patel P. H. The Efficiency of Silver Nanoparticles Synthesized Using Streptomyces spp. Against Human Pathogens. Biosc.Biotech.Res.Comm. 2021;14(2).
Kulshrestha A. K, Patel P. H. The Efficiency of Silver Nanoparticles Synthesized Using Streptomyces spp. Against Human Pathogens. Biosc.Biotech.Res.Comm. 2021;14(2). Available from: <a href=”https://bit.ly/3eIFf8t“>https://bit.ly/3eIFf8t</a>
Copyright © Kulshrestha and Patel This is an open access article distributed under the terms of the Creative Commons Attribution License (CC-BY) https://creativecommns.org/licenses/by/4.0/, which permits unrestricted use distribution and reproduction in any medium, provide the original author and source are credited.
INTRODUCTION
Use of silver dates back to 69 B.C.E. The use of silver has been happening for a long time and was reported from Before Christ to mentioned year (Hill et al. 1939; Alexander 2009). However chemical reduction has been the major technique, production of nanoparticles is possible with various life forms (Mohanpuria et al. 2008; Kholoud et al. 2010). NADH co-factor of nitrate reductase plays a role in the reduction process (Husseiny et al. 2007). AgNO3 concentration, pH, and reaction temperature are decisive in obtaining the silver nanoparticles of desired size (Gurunathan et al. 2009). Bio-synthesized nanoparticles should not only be effective against pathogens but also be nontoxic to the patient and this property is possessed by silver nanoparticles (Song et al. 2009).
Pseudomonas stutzeri can produce AgNPs in its periplasmic space, whereas Verticillium has the ability to produce them on the surface of mycelia as its surface contains nitrate reductase, in turn, this enzyme can bind silver ions with its negatively charged groups and gives the output as reduced silver ions (Klaus et al. 1999; Senapati et al. 2004; Krishnamurthy et al. 2010). Silver nanoparticles can accumulate inside the bacterial cells and kill them, in turn; these killed cells can act as reservoirs of AgNPs and release silver cations slowly in the environment of pathogenic bacteria (Mohamed et al. 2020).
Microbes arthrospira commonly used as dietary supplements have also been found to have the ability with some biochemical alterations during the process to produce nanoparticles (Cepoi et al. 2015). AgNPs are potential agents for treating viruses, cancers of breast, liver, skin and blood (Wei et al. 2015). High doses of antibiotics have side effects, if silver nanoparticles added to antibiotics cause the reduction of antibiotics’ dose with the same or increased efficiency as compared to the previous high dose of antibiotics’ alone, harmful effects of high doses can be minimized (Singh et al. 2015).
Not only the enzymes like nitrate reductase but also the substances having carboxyl, amino, or hydroxyl groups have been noticed to reduce and cap the ions (Abdel-Raouf et al. 2017). Due to their smaller size, Silver nanoparticles can cross the membranes of pathogens and cause the production of free radicals (Siddiqi et al. 2018). Since silver nanoparticles in high doses can cause the leakage of hemoglobin from erythrocytes hence exhaustive examination of the In Vivo effects must be followed (Hamouda et al. 2019). This work presents a comparative account of lethal effects of the strain synthesized AgNPs and antibiotics against human pathogens.
MATERIAL AND METHODS
The soil sample was collected at the bank of the heavily polluted Tapi River, Surat (Gujarat). The sample was sprinkled on actinomycetes isolation agar media, four strains naming AK1-AK4 were isolated, transferred to starch casein broth in Erlenmeyer flasks and grown at 120 rpm, and 300 c for 72 hours. Preliminary identification of these strains was done with citrate utilization, methyl red, indole, urea hydrolysis, and nitrate reduction tests. Broths of flasks; AK1-AK4 in two sets, were filtered through bacteriological filter paper, and two types of filtrates; cell filtrate and cell-free filtrate, were obtained.
One set was left as control. 10ml of each filtrate; AK1-AK4, from one set was made up to 48.5 ml with phosphate buffer of 0.1M and then 1ml of 1mM AgNO3 each and 0.5 ml of 1mM of methionine and cysteine each were added to these filtrates. The same was followed with a set of controls except the addition of AgNO3. Flasks of these filtrates were kept in a shaking incubator at 120 rpm and 300 C for 72 hours and any colour change to brown; an indication of silver nanoparticles’ synthesis, was noticed (Sukanya et al. 2013).
This 50 ml was evaporated to have 5 ml as a semi-final volume and a further 10 times dilution of this semi-final volume for readings with spectrophotometer was done leaving one set as undiluted semi-final volume for the highly resistant bacteria. Separation of antibiotics was done with a dialysis membrane and phosphate buffer of 6.5pH as silver nanoparticles are conjugated with bio-polymers and cannot pass through the pores of the membrane. Spectrophotometric analysis was done at 420 nm. On finding the maximum absorbance given by the product of strain AK3, the candidate was got sequenced for 16S rRNA with ABI 3130 genetic analyzer at Biokart Ltd. Bangaluru and zeta potential and size distribution of silver nanoparticles synthesized by Streptomyces spp. AK3 (Gen Bank MT626067) were measured and checked for the degree of agglomeration at PERD, Ahmedabad.
Strain AK3 was analyzed for shape and size with TEM at Sprint testing, Powai, Mumbai after some improvement in procedure with the addition of amines to prevent the agglomeration. Enterococcus foecails, Coagulase negative Staphylococci spp., Klebsiella pneumoniae, Staphylococcus aureus and, Salmonella spp. were selected for testing the silver nanoparticles’ activity. These microbes were grown on Petri plates with different concentrations of synthesized silver nanoparticles; given in table-1, and antibiotics in the wells.
RESULTS AND DISCUSSION
Morphological and physiological analyses: Colonies were wrinkled, raised, opaque and, white and strains were visualized as long positive rods. Strains could grow in the range between 260C to 400C and the best growth was observed at 320C.Production of AgNPs was checked at pH ranging from 4.5 to 8.5. The optimum pH for production was found as 6.5. Slightly acidic pH may impart a stronger binding ability to capping agents, in turn; smaller sized nanoparticles are formed (Gan et al. 2012). 2% NaCl was the optimum concentration at which strain can grow the best as opposed to halophytic actinomycetes which grow at 15%-25% salt concentration (Jiang et al. 2016).
Biochemical tests: The strain showed positive results for citrate utilization, methyl red, urea hydrolysis and nitrate reduction, and negative for indole.
UV-Visible spectroscopic analysis of synthesized nanoparticles:
Figure 1: Absorbance on Y axis at wavelengths in nm
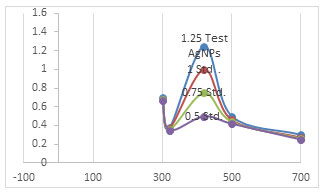
Figure of absorbance versus wavelength was developed with Shimadzu UV-1601 spectrophotometer. An inference can be made for the synthesis of silver nanoparticles with the colour change to light brown. The test sample was diluted ten times and readings were noted with standards of 40 μg/ml, 30 μg/ml and 20 μg/ml as 1.25, 1, 0.75, and 0.5 absorbances respectively from the test sample to the least concentrated standard. The test sample was diluted later also as per the matches with the most effective antibiotics available in terms of efficiencies against human pathogens, whose data are described in this document. Surface plasma resonance of AgNPs synthesized by Streptomyces spp. ranges from 440-450 nm (Eid et al. 2020).
16S rRNA sequencing of strain AK3: Strain AK3 (GenBank MT626067) was found as Streptomyces spp. closer to Streptomyces atacamensis. A few species of Streptomyces are known to have the ability to convert silver ions to silver nanoparticles. Primers such as 27 F and 1492 R can be used for the amplification of Streptomyces spp. (Khadyat et al. 2020).
Zeta potential silver nanoparticles: Zeta potential was-14.4 which shows agglomeration-free silver nanoparticles to the most extent. AgNPs can be sterically stabilized by natural surfactants (Chartarrayawadee et al. 2020).
Figure 2: Synthesized silver nanoparticles’ Zeta potential measured by Zeta sizer

Features of produced silver nanoparticles known with TEM:
Figure 3: Images of silver nanoparticles with TEM
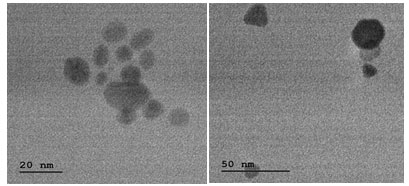

TEM produced the images with 20 nm-sized silver nanoparticles having various morphological characteristics such as spherical, prism-shaped etc. A role against destabilization of silver nanoparticles is played by sulphur-containing amino acids, thiols amines, alcohols to some extent as different results are seen for control and stabilizers added silver nanoparticles synthesizing cell-free filtrate (Iravani et al. 2014).
Properties can be manipulated by changing the concentration of capping agents, however the efficiency of silver nanoparticles against pathogens is affected with increased concentration adversely. Antimicrobial activity of silver nanoparticles depends on the shapes also. Dispersity of AgNPs can be modulated by the flow rate in a continuous flow tubular microreactor (Dawadi et al. 2021).
Comparative antibacterial activities of synthesized silver nanoparticles and antibiotics: Well diffusion method was performed and data of affected microbes by different antibiotics and SNPs with zones of inhibition have been represented here as Petri plates and the table is shown as well below them.
Figure 4: Inhibition zones of SNPs and antibiotics
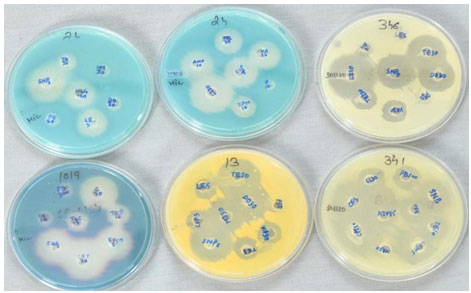
24 (a)- Enterococcus foecalis with SNPs, LE5, and HIG-120
24 (b)– Enterococcus Foecalis with LZ30, AMP10, TEI30, IPM10, VA30, and CIP5
346-Co-agulase negative Staphylococci spp. with SNPs, LZ30, TEI30, VA30, DO30, and TE30
1019-Klebsiella pneumoniae with SNPs, CPT30, CFM5, C30, TOB10, TFC15
13-Staphylococcus aureus with SNPs, TE30, LE5, CIP5, TEI30, DO30, and MI30
341-Salmonella spp. with SNPs, PB300, CL10, CFS30, TE10, CFM5, AZM15, and PF5
Table 1. Effect of Streptomyces’s strain AK3synthesized Silver Nanoparticles (SNPs) on test organisms
| Test Organism | Antibiotics-μg/ SNPs | Zone of inhibition (mm) | Plate No. |
| Staphylococcus aureus | Tetracycline 30 (TE30) | 28 | 13 |
| SNPs | 25 | ||
| Levofloxacin 5 (LE5) | 12 | ||
| Ciprofloxacin 5 (CIP5) | 9 | ||
| Penicillin-G 10 (P10) | 7 | ||
| Vancomycin 30 (VA30) | 15 | ||
| Teicoplanin 30 (TEI 30) | 14 | ||
| Doxicycline 30 (DO 30) | 13 | ||
| Minocycline 30 (MI 30) | 23 | ||
| Clindamycin 2 (CD2) | 18 | ||
| Salmonella spp. | SNPs | 22 | 341 |
| Polymyxin B 300 (PB300) | 15 | ||
| Colistin 10 (CL10) | 17 | ||
| Cefopazone sulbactum30 (CFS30) | 24 | ||
| Tetracline 10 (TE10) | 17 | ||
| Cefixime 5 (CFM5) | 25 | ||
| Azithromycin 15 (AZM15) | 16 | ||
| Pefloxacin 5 (PF5) | 18 | ||
| Coagulase negative Staphylococci | Linezolid 30 (LZ30) | 26 | 346 |
| SNPs | 24 | ||
| Teicoplanin 30 (TEI30) | 15 | ||
| Vancomycin 30 (VA30) | 16 | ||
| Doxycycline 30 (DO30) | 20 | ||
| Tetracycline 30 (TE30) | 23 | ||
| Table 1 Cont. | |||
| Test Organism | Antibiotics-μg/ SNPs | Zone of inhibition (mm) | Plate No. |
| Enterococcus faecalis | Linezolid 30 (LZ30) | 25 | 24 (a) |
| Ampicillin10 (AMP10) | 19 | ||
| Teicoplanin 30 (TEI30) | 17 | ||
| Imipenam 10 (IPM10) | 15 | ||
| Vancomycin 30 (VA30) | 19 | ||
| Ciprofloxacin 5 (CIP) | 20 | ||
| Enterococcus faecalis | SNPs | 24 | 24 (b) |
| Levofloxacin 5 (LE5) | 18 | ||
| High level gentamycin 120 (HIG120) | 20 | ||
| Klebsiella pneumoniae | SNPs | 25 | 1019 |
| Ceftaroline 30 (CPT30) | 27 | ||
| Cefixime 5 (CFM5) | 22 | ||
| Chloramphenicol 30 (C30) | 22 | ||
| Tobramycin 10 (TOB10) | 15 | ||
| Tigecycline 15 (TGC15) | 13 | ||
Staphylococcus aureus, Salmonella typhi, Pseudomonas aeruginosa, E. coli, Co-agulase negative Staphylococci, Enterococcus foecalis, and Klebsiella pneumoniae were selected as test pathogens. 2. Staphylococcus aureus, Salmonella typhi, Pseudomonas aeruginosa, E. coli, and Co-agulase negative Staphylococci were tested on Mueller Hinton agar whereas Enterococcus foecalis and Klebsiella pneumoniae were tested on Hi-chrome media.
Plates were placed in the incubator for 20 hours at 370 C. 3. Staphylococcus aureus was highly sensitive to tetracycline 30, almost resistant to penicillin-G 10, and sensitive to silver nanoparticles (10 μg/ml) with a 25 mm zone of inhibition. Addition of silver nanoparticles to the tetracycline, produced in the current work, could have enhanced lethal effect on S. aureus (Hussein et al. 2019). Effect of silver nanoparticles on the closer species Staphylococcus epidermidis has been reported (Swolana et al. 2020)
Test organism Salmonella was the most sensitive to cefapazone sulbactam 30, the least sensitive to polymyxin B300 and sensitive to silver nanoparticles with a 22 mm inhibition zone. The sensitivity of Salmonella to silver nanoparticles (10 μg/ml) has been documented as the “sensitivity depends on the strain, and dose” (Losasso et al. 2014; Petrus et al. 2011). When Salmonella braenderup was exposed to AgNPs, membranes of the microbes were ruptured (Diego et al. 2020).
Co-agulase negative staphylococci can cause infections when they reach the blood stream. An iinhibition zone of 25 mm was produced by prepared AgNPs. Linezolid was the only antibiotics with a bigger zone of 26 mm than that of AgNPs.
Treatment of Enterococcus foecalis is difficult when it reaches the urinary tract (Kau et al. 2005). Linezolid 30 is effective against this species. In the current study, Linezolid 30 produced a 25 mm of zone of inhibition. Prepared AgNPs (30μg/ml) had almost the same effect as that of Linezolid with a 24 mm zone of inhibition. Traditional approaches are less effective in treatment of Enterococcus foecalis however; silver nanoparticles could be a potent solution to treat them (Sadony et al. 2019).
Klebsiella pneumoniae produces extended spectrum β- lactamase, in turn, this compound can breakdown many medicines. Carpabenam has been found effective against ESBL-producing Klebsiella pneumonia (Patersonet al. 2004). However, Klebsiella pneumoniae shows greater resistance against considerably safe AgNPs but at higher doses as 100μg/ml, AgNPs formed a 25 mm zone of inhibition. Klebsiella pneumoniae has been resistant to ampicillin and AgNPs were tested against this species to get effective results (Hamida et al. 2020).
CONCLUSION
Streptomyces Spp. AK3 has been found to synthesize expected quality silver nanoparticles. Synthesized silver nanoparticles with this strain are much closer to the most effective antibiotics against human pathogens in terms of the formation of zones of inhibition as represented in table-1. The strain synthesized silver nanoparticles should be preferred to antibiotics for the sake of having lesser side effects, being cost-effective, and working as a tool against multi-drug resistant bacteria.
CONCLUSION
Streptomyces Spp. AK3 has been found to synthesize expected quality silver nanoparticles. Synthesized silver nanoparticles with this strain are much closer to the most effective antibiotics against human pathogens in terms of the formation of zones of inhibition as represented in table-1. The strain synthesized silver nanoparticles should be preferred to antibiotics for the sake of having lesser side effects, being cost-effective, and working as a tool against multi-drug resistant bacteria.
ACKNOWLEDGEMENTS
The work was supported by PERD, Ahmedabad, Biokart Ltd., Bangaluru and Sprint testing solutions, Mumbai.
Conflict of interests: Authors declare no competing interests among themselves.
REFERENCES
Abdel-Raouf, N., Al-Enazi, N.M., and Ibraheem, I.B.M., (2017), Green biosynthesis of gold nanoparticles using Galaxaura elongata and characterization of their antibacterial activity, Arabian Journal of Chemistry, Vol. 10, S3029-S3039.
Abou El-Nour Kholoud, M.M., Eftaiha, Ala’a, Al-Warthan, and A., Ammar Redda A.A., (2010), Synthesis and applications of silver nanoparticles, Arabian Journal of Chemistry, Vol. 3, Pages 135-140.
Alexander, J.W., (2009), History of the medical use of the silver, Surgical Infections,Vol. 10, No.3, Pages 289-292.
Cepoi, L., Chiriac, T., Valuta, A., Zinicovscaia, I., Kirkesali, E., Frontasyeva, M., Culicov, O., Pavlov, S., and Bobrikov I., (2015), Biochemical changes in cyanobacteria during the synthesis of silver nanoparticles, Canadian Journal of Microbiology, Vol. 61, No. 1, Pages 13-21.
Chartarrayawadee, W., Charoensin, P., Saenma, J., Rin, T., Khamai, P., Nasomjai, P., and Too, C.O., (2020), Green synthesis and stabilization of silver nanoparticles using Lysimachia foenum-graecum hance extract and their antibacterial activity, De Gruyter, DOI: https://doi.org/10.1515/gps-2020-0012.
Dawadi, S., Katuwal, S., Gupta, A., Lamichhane, U., Thapa, R., Jaisi, S., Lamichhane, G., Bhattarai, D.P., and Parajuli, N., (2021), Current research on silver nanoparticles: synthesis, characterization, and applications, Journal of nanomaterials, Vol. 2021, Article ID 6687290.
Diego, V., Liliana, C., Tatiana, H., Felipe, C., Graciela, J., and Brayan, V., (2020), Effect of silver nanoparticles on the morphology of Toxoplasm gondii and Salmonella braenderup, Journal of nanotechnology, Vol. 2020, Article ID 9483428.
Eid, A.M., Fouda, A., Niedbala, G., Hassan, S.E., Salem, S.S., Abdo, A.M., Hetta, H.F., and Shaheen, T.I., (2020), Endophytic Stretomyces larentii mediated green synthesis of AgNPs with antibacterial and anticancer properties for developing functional textile fabric properties, Antibiotics, 9, 641.
Gan, P.P., Ng, S.H., Huang, Y., and Li, S.F.Y., (2012), Green synthesis of gold nanoparticles using palm oil mill effluent (POME): a low-cost and eco-friendly viable approach, Bioresource Technology, Vol. 113, Pages 132-135.
Gurunathan, S., Kalishwaralal, K., Vaidyanatha, R., Vekataraman, D., Pandian, S., Muniyandi, J., Hariharan, N., and Eom, S.H., (2009), Biosynthesis, purification and characterization of silver nanoparticles using E. coli, Vol. 74, Issue 1, Pages 328-335.
Hamida, R.S., Ali, M.A., Goda, D.A., Khalil, M.I., and Redhwan, A., (2020), Cytotoxic effect of green silver nanoparticles against ampicillin-resistant Klebsiella pneumonia, RSC advances, 10, 211336-21146.
Hamouda, R.A., Hussein, M.H., Abo-elmagd, R.A., (2019), Synthesis and biological characterization of silver nanoparticles derived from the Cyanobacterium Oscillatoria limnetica, Scientific Reports, Vol. 9, and Article No. 13071.
Hill, W.R., and Pillsbury, D.M., (1939), Argyria; The Pharmacology of silver, Baltimore. The Williams and Wilkins Company.
Hussein E.A.M., Mohammad A.A., Harraz F.A., Ashan M.F., (2019), Biologically synthesized silver nanoparticles for enhancing tetracycline activity against Staphylococcus aureus and Klebsiella pneumoniae, Brazilian archives of biology and technology, vol. 62.
Husseiny, M., El-Aziz, M.A., Badr, Y., and Mahmoud M.A., (2007), Biosynthesis of gold nanoparticles using Pseudomonas aeruginosa, Spectrochimica Acta part-A: Molecular and Biomolecular Spectroscopy, Vol. 67, No. 3-4, Pages 1003-1006.
Iravani, S., Korbekandi, H., Mirmohammadi, S.V., and Zolfaghari B., (2014), Synthesis of silver nanoparticles: chemical, physical and biological methods, Research in Pharmaceutical Sciences, Vol. 9, Issue 6, Pages 385-406.
Jiang, Y., Li Q., Chen X., and Jiang C., (2016), Actinobacteria: basics and biotechnological application, DOI-10.5772/61457.
Kau, A.L., Martin, S.M., Lyon, W., Hayes, E., Caparon, M.G., and Hultgren S.J., (2005), Enterococcus foecalis tropism for the kidneys in urinary tract of C57BL/6J mice, (2005), Infection and Immunity, Vol.73, Issue 4, Pages 2461-2468.
Khadyat, K., Shepra, D.D., Malla, K.P., Shrestha, S., Rana, N., Marasini, B.P., Kanal, S., Rayamajhee, B., Bhattarai, B.R., and Parajuli, N., (2020), Molecular identification and antimicrobial potential Streptomyces species from Nepalese soil, International journal of microbiology, Vol. 2020, Article ID 881747.
Klaus, T., Joerger, R., Olsson, E., and Granqvist C-G., (1999), Silver-based crystalline nanoparticles, microbially fabricated, Proceedings of the National Academy of Sciences of the United State of America, Vol. 96, No. 24, Pages 13611-13614.
Krishnamurthy, S., Mao, J., Satishkumar, M., Kwak, I.S., and Yun, Y.S., (2010), Corynebacterium glutanicum mediated crystallization of silver ions through sorption and reduction process, Chemical Engineering Journal, Vol. 162, No. 3, Pages 989-996.
Losasso, C., Belluco, S., Cibin, V., Zavagnin, P., Micetic, I., Gallocchio, F., Zanella, M., Bregoli, L., Biancotto, G., and Ricci A., (2014), Antibacterial activity of silver nanoparticles: sensitivity of different Salmonella serovars, Frontiers in Microbiology, Vol. 5, Pages 227.
Mohamed, D.S., El-Baky, R.M.A., Sandle, T., Mandour, S.A., and Ahmed E.F., (2020), Antimicrobial activity of Silver-Treated Bacteria against other Multi-Drug Resistant Pathogens in Their Environment, Antibiotics, Vol. 9, Issue 4, Article No. 181.
Mohanpuria, P., Rana, N.K., and Yadav, S.K., (2008), Biosynthesis of nanoparticles: technological concepts and future applications, Journal of Nanoparticle Research, Vol. 10, Issue 3, Pages 507-517.
Petrus, E.M., Tinakumari, S., Chai, L.C., Ubong, A., Tunung, R., Elexson, N., Chai, L.F., and Son R., (2011), A study on minimum inhibitory concentration and minimum bactericidal concentration of nano colloidal silver on food borne pathogens, International Food Research Journal, Vol. 18, Issue 1, Pages 55-66.
Paterson, D.L., Ko, W-C., Gottberg, A.V., Mohapatra. S., Casllas, J.M., Goossens, H., Mulazimoglu, L., Trenholme, G., Klugman, K.P., Bonomo, R.A., Rice, L.B., Wagener, M.M., MacCormack, J.G., and Yu, V.L., (2004), Antibiotic therapy for Klebsiella pneumoniae bacteremia: Implications of production of extended spectrum β- Lactamases, Clinical Infectious Diseases, Vol. 39, Issue 1, Pages 31-37.
Sadony, D.M., and Montasser, K., (2019) Evaluation and comparison between the bactericidal effect of diode laser irradiation (970 nm) and silver nanoparticles on Enterococcus foecalis bacterial strain (as in vitro study), Bulletin of national research centre, 43, 155, doi.org/10.1186/s42269-019-0188-5.
Senapati, S., Mandal, D., Ahemad, A., Khan, M.I., Sastry, M., and Kumar, R., (2004), Fungus mediated synthesis of silver nanoparticles: A novel biological approach, Indian Journal of Physics, Vol. 78, No. 1, Pages 101-105.
Siddiqi, K.S., Husen, A., Rao, and Rifaqat A.K., (2018), A review on biosynthesis of silver nanoparticles and their biocidal properties, Journal of Nanobiotechnology, Vol. 10, Art. No.14.
Singh, P., Kim, Y., Singh, H., Wang, C., Hwang, K.H., Yang, D.C., and Farh, M.E., (2015), Biosynthesis, characterization, and antimicrobial applications of silver nanoparticles, International Journal of Nanomedicine, Vol. 10, Issue 1, Pages 2567-2577.
Song, J.Y., and Kim, B.S., (2009), Rapid biological synthesis of silver nanoparticles using plant leaf extracts, Bioprocess and Biosystems Engineering, Vol. 32, Issue 1, Pages 79-84.
Sukanya M.K., Saju K.A., Praseetha P.K., Sakthivel G., (2013), Therapeutic potential of biologically reduced silver nanoparticles from actenomycete culture, Journal of Nanoscience, Vol. 2013, Article ID 940719.
Swolana, D., Kepa, M., Idzik, D., Dziedzic, A., Kabala-Dzik, A., Wasik, T.Z., and Wojtczka, R.D., (2020), The antibacterial effect of silver nanoparticles on Staphylococcus epidermidis strains with different biofilm—forming ability, Nanomaterials, Vol. 10, Issue 5, https://www.mdpi.com/2079-4991/10/5/1010.
Wei, L., Lu, Z., Xu, H., Patel, A., Chen, Z-S., and Chen, G., (2015), Silver nanoparticles: synthesis, properties and therapeutic applications, Drug Discovery Today, Vol. 20, Issue 5, Pages 595-601.


