Cellulase production in
Lysinibacillus
sp isolated from
the estuaries of Odisha
Shubhashree Mahalik,
1
* Deepali Mohapatra
2
and Dhanesh Kumar
3
1,2
Post Graduate Department of Biosciences and Biotechnology, Fakir Mohan University, New campus,
Nuapadhi, Balasore, Odisha-756020, India
3
School of Life Sciences, Hyderabad Central University, Prof C. R. Rao Road, P.O. Central University,
Gachibowli, Hyderabad, Telangana 500046, India
ABSTRACT
Microbes are rich sources of natural products like secondary metabolites, enzymes and proteins. In this context the
primary objective of the present work is to isolate microbes from natural habitats and characterize them on the basis
of their ability to produce metabolites. Since estuaries are the junction of marine and coastal habitats and harbour a
plethora of microbes therefore in this study estuaries along the coastal district of Balasore, Odisha were screened for
cellulase secreting bacteria. Cellulase forms an important component of enzyme cocktail used for degradation of lig-
nocellulosic biomass for production of biofuel. Several rounds of sampling, pure culture, morphological, biochemical
and phylogenetic screening led to the identi cation of Lysinibacillus sp. having the ability to secrete cellulase. Physi-
cal as well as nutritional characterization like optimization of media, temperature, carbon and nitrogen requirements
was performed to enhance the biomass formation. The isolated strain of Lysinibacillus sp. showed higher biomass and
growth rate at 37°C, in Terri c Broth media supplemented with 0.5% Glucose and 0.5% Sodium Nitrite. Submerged
fermentation under anaerobic condition at 37°C for 5 days led to release of 9.85µmole of glucose/ml of enzyme.
KEY WORDS:
LYSINIBACILLUS
, ESTUARY, CELLULASE, PHYLOGENETIC ANALYSIS
743
Biotechnological
Communication
Biosci. Biotech. Res. Comm. 11(4): 743-753 (2018)
ARTICLE INFORMATION:
Corresponding Authors: shubhashreemahalik@gmail.com
Received 21
st
Oct, 2018
Accepted after revision 21
st
Dec, 2018
BBRC Print ISSN: 0974-6455
Online ISSN: 2321-4007 CODEN: USA BBRCBA
Thomson Reuters ISI ESC / Clarivate Analytics USA
Mono of Clarivate Analytics and Crossref Indexed
Journal Mono of CR
NAAS Journal Score 2018: 4.31 SJIF 2017: 4.196
© A Society of Science and Nature Publication, Bhopal India
2018. All rights reserved.
Online Contents Available at: http//www.bbrc.in/
DOI: 10.21786/bbrc/11.4/27

Shubhashree Mahalik et al.
744 CELLULASE PRODUCTION IN
LYSINIBACILLUS
SP ISOLATED FROM THE ESTUARIES OF ODISHA BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS
INTRODUCTION
Several microorganisms are considered to be extremo-
philes as they are adapted to extreme environmental
conditions such as high or low temperature, alkaline
or acidic water, high pressure and substrate limitations.
These characters make them potential industrial organ-
isms as they produce several interesting metabolites to
cope these extreme conditions. Various literatures are
available where microorganisms isolated from extreme
geographical locations have been used for production
of hormones, proteins, enzymes and other primary and
secondary metabolites (Coker, 2016; L Bergquist et al.,
2014; Littlechild, 2015; Poli et al., 2017; Stierle & Stierle,
2014; Yin & Chen, 2015). One such extreme environ-
ment is the estuary. Estuaries have rich biodiversity due
to variation in temperature, pH, salinity and availability
of salts and other minerals (Campbell & Kirchman, 2013;
Lallias et al., 2015). Several Bacillus species are known
to be cellulase producers (Irfan et al., 2017; Sanjeev et
al., 2017), but very few reports are available on cellulase
production by Lysinibacillus strains
These brackish water bodies are an amalgamation of
both fresh water from rivers and saline water from tidal
waves of sea. This character makes it a very productive
habitat for various ora and fauna including microor-
ganisms (Moyle et al., 2010). Odisha is one of the coastal
states situated on the eastern part of India having a
coast line of 480 km. The coastline of the Balasore dis-
trict is in the shape of a strip with a length of 81 km and
26 km wide. Several estuarine rivers like Budhabalanga,
Subarnarekha ood the coastal areas. Many studies have
been carried out on the macro ora and fauna of this
area but very few documented information is available
regarding the microbial biodiversity, (Subudhi & Patra,
2013; Sujana et al., 2015 Bomble et al., 2017).
Cellulases are a group of enzyme that degrade cel-
lulose by hydrolyzing -1,4 linkages in cellulose chains.
Naturally, the cellulase is produced from widespread
sources like fungi, bacteria, protozoans, plants, and
animals (symbiotic bacteria in few ruminants and ter-
mites). The biotechnological application of lignocel-
luosic biomass in several industries like paper, textile,
food, biofuel as well as agriculture has led to extensive
research on production, biochemical as well as enzy-
matic characterization of cellulase. Microbes from many
bacterial genera have proved to be a good source for
cellulase production at industrial scale, Juturu & Wu,
2014; Kuhad, Gupta, & Singh, 2011 Kuhad et al., 2016
and Bomble et al., 2017). In the present study an attempt
has been made to isolate cellulase producing bacte-
ria from the estuaries around the Balasore district and
optimize the culture conditions for the production of
cellulase.
MATERIALS & METHODS
Sample collection and pure culture: Soil sediments up to
a depth of 6-10 inch were collected from different spots
of the Khandia estuary (21°19’1.65”N; 86°53’32.99”E) in
Balasore district near the mouth of Khandia river. Soil
samples were serially diluted in 1X PBS solution and
plated on nutrient agar (NA) plate. The plates were incu-
bated at 37°C for 24 hour and the colonies obtained were
re-streaked on fresh NA plate and the mother plate was
allowed to incubate further. The colonies that appeared
after 48 hour and 72 hour of incubation were also re-
streaked on fresh NA plates to obtain isolated colonies.
Dilution streaking was performed for all the isolated
colonies that were obtained from various samples. This
process was repeated for several times till pure cultures
were obtained. All pure cultures were labelled and stored
at 4°C in NA stabs.
Screening for cellulase producing strains: The
isolates were screened on the basis of their ability to
secrete cellulase. NA plates were overlayed with 0.5%
carboxymethyl cellulose (CMC). CMC acts as substrate
for cellulase enzyme. The bacterial cultures were spread
on the CMC supplemented plates. Post incubation the
plates were stained with 0.1% Congo Red. The plates
were destained with 1N NaCl solution and observed for
clear zones (Meddeb-Mouelhi, Moisan, & Beauregard,
2014).
Morphological and physiological characterization of
the isolates: The shape, size, elevation, margin and col-
our of the colony were observed and the morphology
of the isolates was determined using Grams Staining
method. Biochemical test like Citrate Utilization, Triple
Sugar Iron, Mannitol Motility, Gelatin Hydrolysis, Oxi-
dase, Indole production and antibiotic resistance tests
were performed. Citrate Utilization was performed on
Simmon’s citrate agar medium (Himedia, M099), TSI
was tested on Triple Sugar-Iron Agar (Himedia, MM021),
Mannitol Motility was checked on Mannitol Motility
Test Medium (Himedia, M770), Gelatin Hydrolysis was
checked on Nutrient gelatin medium prepared in lab.
The presence of oxidase enzyme was checked using
Oxidase disc (Himedia, DD018) and Indole production
was checked using Kovac’s strip (Himedia, DD019). All
experiments were performed using standard protocol as
recommended by product manual. Resistances for ampi-
cillin, kanamycin, tetracycline, penicillin and strepto-
mycin were checked by disc diffusion method at 50g/
l, 5g/l, 0.5g/l and 0.05g/l concentration for all
the antibiotics.
Growth Characteristics of the Isolates: The isolates
were grown in 4 different media (Nutrient Broth, Terri c
Broth, Marine Broth and Arti cial Sea Water) at 25 ºC

Shubhashree Mahalik et al.
BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS CELLULASE PRODUCTION IN
LYSINIBACILLUS
SP ISOLATED FROM THE ESTUARIES OF ODISHA 745
and 37ºC. The experiment was performed in a microbio-
reactor (m2p labs BioLector) where biomass was contin-
uously monitored till the population reached at station-
ary phase. The growth was also monitored under various
carbon sources (Glucose, Maltose, Lactose, Xylose, Cel-
lobiose and starch) and nitrogen sources (Sodium nitrate,
Di-ammonium hydrogen citrate, Ammonium nitrite,
Tryptone, Yeast extract and Ammonium chloride). The
concentration of carbon and nitrogen supplemented to
the media were 0.5% each. Similarly salt tolerance was
checked for the isolates in TB medium supplemented
with various concentration of NaCl ranging from 0.5%
to 12%.
Isolation of Genomic DNA: 5ml bacterial culture was
inoculated and incubated overnight. After incubation
the culture was transferred to a centrifuge tube and cen-
trifuged at 10,000 rpm for 10 minutes until a compact
pellet was formed. The supernatant was discarded and
the pellet was resuspended in a mixture of 567l TE
buffer and 5l RNAse A by repeated pipetting. 15l 10%
SDS and 4l proteinase K (18 mg/mL) was added. It was
mixed thoroughly and incubated 15-20 minutes at 65°C
until all the cells are lysed.100l of 5M NaCl was added
and mixed thoroughly. 80l of CTAB/NaCl (10% w/v;
0.7M) solution was added and mixed thoroughly and
incubated for 10min at 65°C. Equal volume of chloro-
form/isoamyl alcohol (0.7-0.8ml) was added and mixed
thoroughly and centrifuged for 5min at 10,000 rpm. 1
volume of isopropanol was added to the supernatant,
shaken and centrifuged. The pellet obtained was washed
with 70% ethanol, dried and dissolved in 50l TE buffer.
PCR and sequencing: The 16s rRNA gene was ampli ed
from the genomic DNA of all the isolates. The prim-
ers used in the study are as follows (Frank et al., 2008;
Karakasidou et al., 2018): BAC27F AGAGTTTGATCCTG-
GCTCAG BAC1492RGGTTACCTTGTTACGACTT: The PCR
was carried out at initial denaturation of95°C/5min,
denaturation of 95°C/30sec, annealing at 42°C/1min,
extension at 72°C/1min 30sec and nal extension at
72°C/5min. This was repeated for 30 cycles.The sequenc-
ing of the puri ed PCR product was done using the
BAC27F forward primer. The work was outsourced form
SciGenom Labs Pvt. Ltd., Kerala, India.
Phylogenetic Analysis: The molecular characterization
of all the isolates were done by 16s rRNA sequencing.
The phylogenetic was prepared using MEGA 7 program
(Sudhir Kumar, Stecher, & Tamura, 2016) with Neighbor
Joining method. Sequence of Lysinibacillus sp. KEI3 was
BLAST in EZ taxon (Chun et al., 2007) and only val-
idly published sequences were taken as references in tree
formation. Boot strap replication was performed 1000
times.
Submerged Fermentation: Bacterial isolates were cul-
tured in Terri c broth medium under submerged fer-
mentation conditions. 50ml TB medium was prepared
in 250 mL ask. The medium was supplemented with
0.5% Glucose and 0.5% Sodium Nitrite. Medium was
sterilized by autoclaving. The asks were incubated in a
shaking incubator at 37±2°C for 5days and then crude
enzyme was extracted by centrifugation at 10,000rpm
for 20min at 4°C. The cell free culture ltrate (CFCF)
was used as crude enzyme to test Cellulase activity. Cel-
lulase activity was measured by DNS assay.
RESULTS
Isolation of Bacterial Colonies: After serial dilution,
plating of soil samples and incubation, more than 50
colonies were obtained. Out of these, 11 pure cultures
were obtained which were screened for their ability to
degrade CMC by plating them on CMC agar plate and
staining with Congo Red. Isolate 3 showed clear zones
on CMC agar plates indicating their ability to secrete
cellulase. Morphological and biochemical characteriza-
tion was performed for Isolate 3 and the observations
are presented in Table 1. The results indicate that the
isolates belong to genus Bacillus.
Phylogenetic Analysis: Genomic DNA isolated from
pure cultures were subjected to PCR ampli cation of the
Table.1. Biochemical characters of Lysinibacillus sp.KEI-3
TEST
Gram’s staining
TSI
Mannitol Motility
Citrate utilization
Gelatin Hydrolysis
Oxidase
Indole
Ampicillin
Kanamycin
Tetracyclin
Penicillin
Streptomycin
KEI
-3
+K/A- - ++ -+++++
(-) represents negative response to the test; (+) represents positive response or susceptibility to the test.
Shubhashree Mahalik et al.
746 CELLULASE PRODUCTION IN
LYSINIBACILLUS
SP ISOLATED FROM THE ESTUARIES OF ODISHA BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS
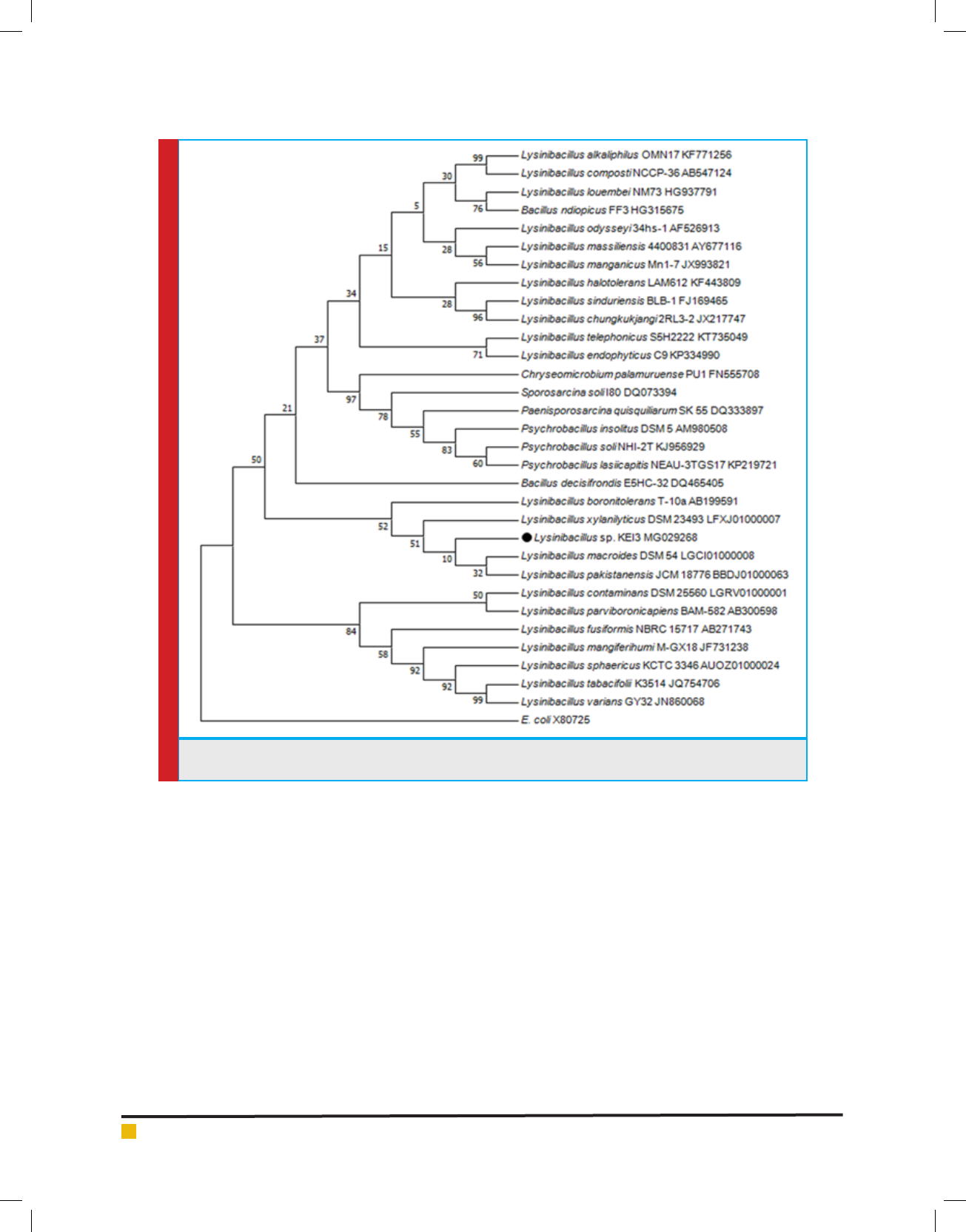
FIGURE 1. Phylogenetic tree prepared using MEGA 7 software with Neighbor Joining methodon the basis of
16sRNA sequencing.
16s RNA using BAC27F and BAC1498R primers which
produced around 1450bp long 16S rRNA gene. Phylo-
genetic tree was prepared using MEGA 7 program and
Isolate 3 was identi ed to be Lysinibacillus sp. which has
maximum similarity to Lysinibacillus fusiformis strain 4
(KF916674) (Figure 1). The Isolate was named as Lysini-
bacillus sp.KEI-3.
Effect of different media on growth
Lysinibacillus sp. KEI-3 was inoculated in Nutrient Broth
(NB), Terri c Broth (TB), Marine Broth (MB) and Arti -
cial Sea Water supplemented with Glucose and Tryptone
(SWGT) media and grown at 25°C and 37°C. The growth
was monitored for 36 and 24 hour respectively and plot-
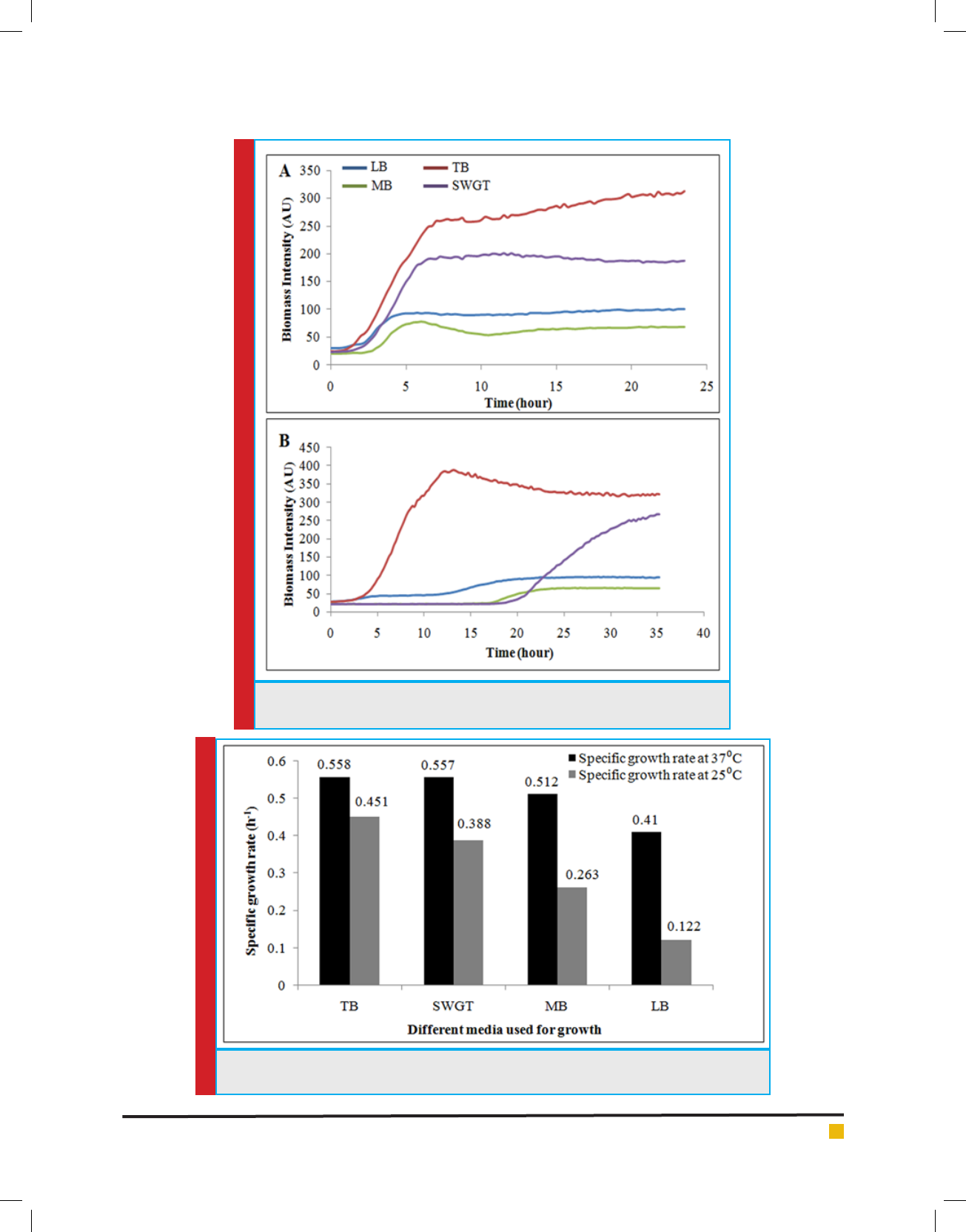
ted to calculate speci c growth rate. It was observed that
when isolates were cultured at 25°C it had a very long
lag phase and the speci c growth rate was also slow in
comparison to cells growing at 37°C (Figure 2). This phe-
nomenon was observed for all media used for the study
except TB medium, where irrespective of temperature
the isolate Lysinibacillus sp.KEI-3 had a high speci c
growth rate. For TB medium the speci c growth rate was
0.558h
-1
at 37°C whereas at 25°C it was 0.451h
-1
which
is almost comparable. Interestingly in SWGT media the
isolate had almost 20 hour long lag phase at 25°C after
which there was increase in growth of the cells and the
maximum speci c growth rate achieved at 25°C was
0.338h
-1
whereas at 37°C the it was 0.557h
-1
(Figure 3).
It was interesting to observe that the isolate could not
grow well in LB and MB. The reasons for LB are obvious
that it is nutritionally less rich and complex than TB due
to which the growth rate was slow. MB is high in salt
Shubhashree Mahalik et al.
BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS CELLULASE PRODUCTION IN
LYSINIBACILLUS
SP ISOLATED FROM THE ESTUARIES OF ODISHA 747
FIGURE 2. Growth pro le of Lysinibacillus sp. KEI-3 strain in different media and
different temperature. (A) growth curve at 37ºC, (B) growth curve at 25ºC.
FIGURE 3. Speci c Growth rate of Lysinibacillus sp. KEI-3 strain in different media and different
temperature.
Shubhashree Mahalik et al.
748 CELLULASE PRODUCTION IN
LYSINIBACILLUS
SP ISOLATED FROM THE ESTUARIES OF ODISHA BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS
concentration and probably the isolate do not have high
salt tolerance, which could be the possible reason for
limited growth in MB medium. SWGT medium was for-
mulated in lab where the composition of arti cial sea-
water was almost similar to the original seawater but the
salt concentration was less. Further it was enriched by
addition of Glucose and Trypotone which are very good
sources of carbon and complex nitrogen requirements.
Effect of different carbon source on growth: Six dif-
ferent carbon sources namely Glucose, Lactose, Starch,
Xylose, Maltose and Cellobiose were selected in present
study. These carbon sources were added at a nal con-
centration of 0.5% separately in TB media. Lysinibacillus
sp.KEI-3 was inoculated in these media and grown at
37°C. The growth was measured after 24 hour of incuba-
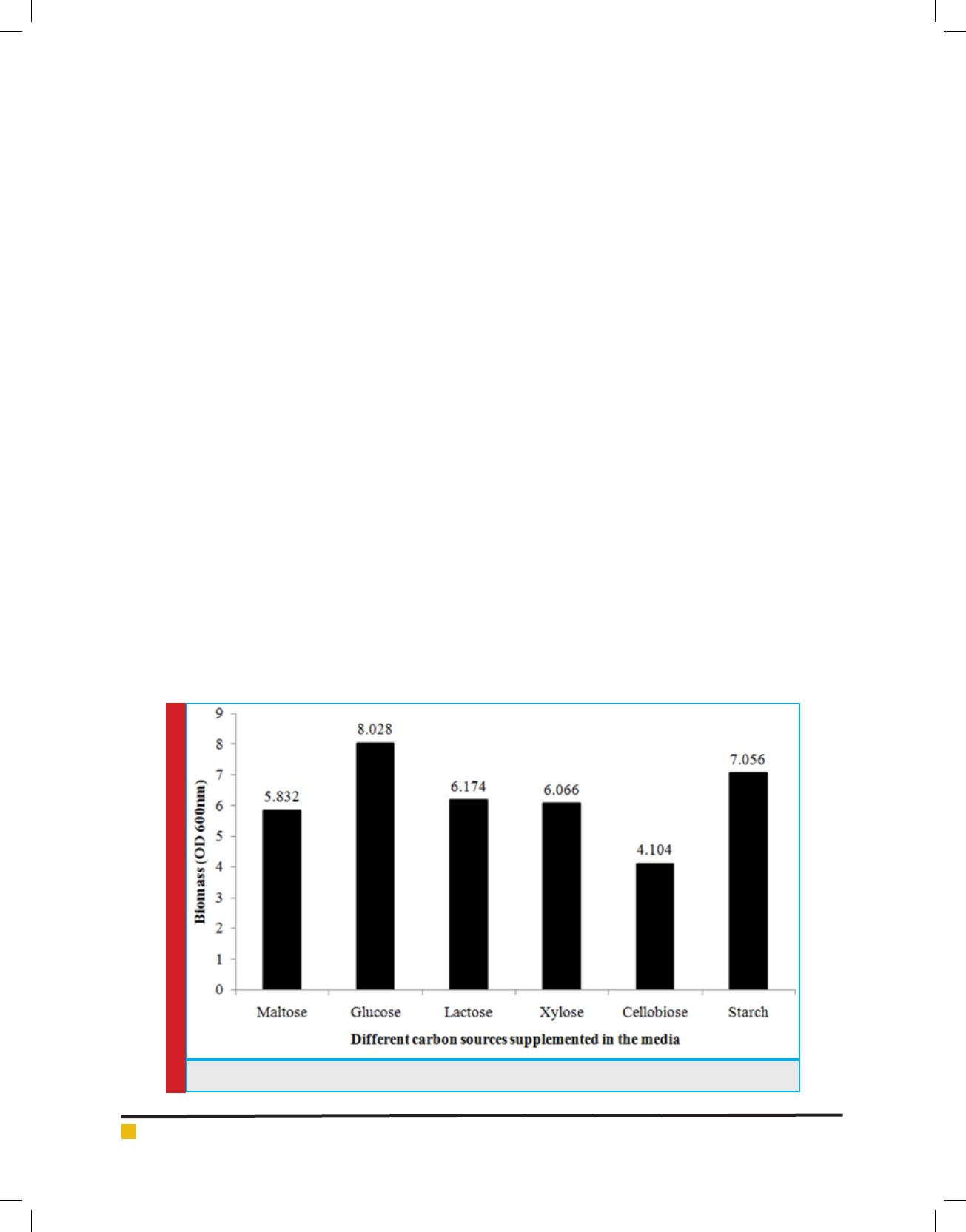
tion. It was observed that the isolate could effectively
utilize both pentose and hexose sugars. Biomass pro l-
ing revealed that Lysinibacillus sp. KEI-3 effectively uti-
lize monosaccharide (glucose and xylose), disaccharides
(maltose, lactose and cellobiose) as well as polysaccha-
ride (starch) (Figure 4).
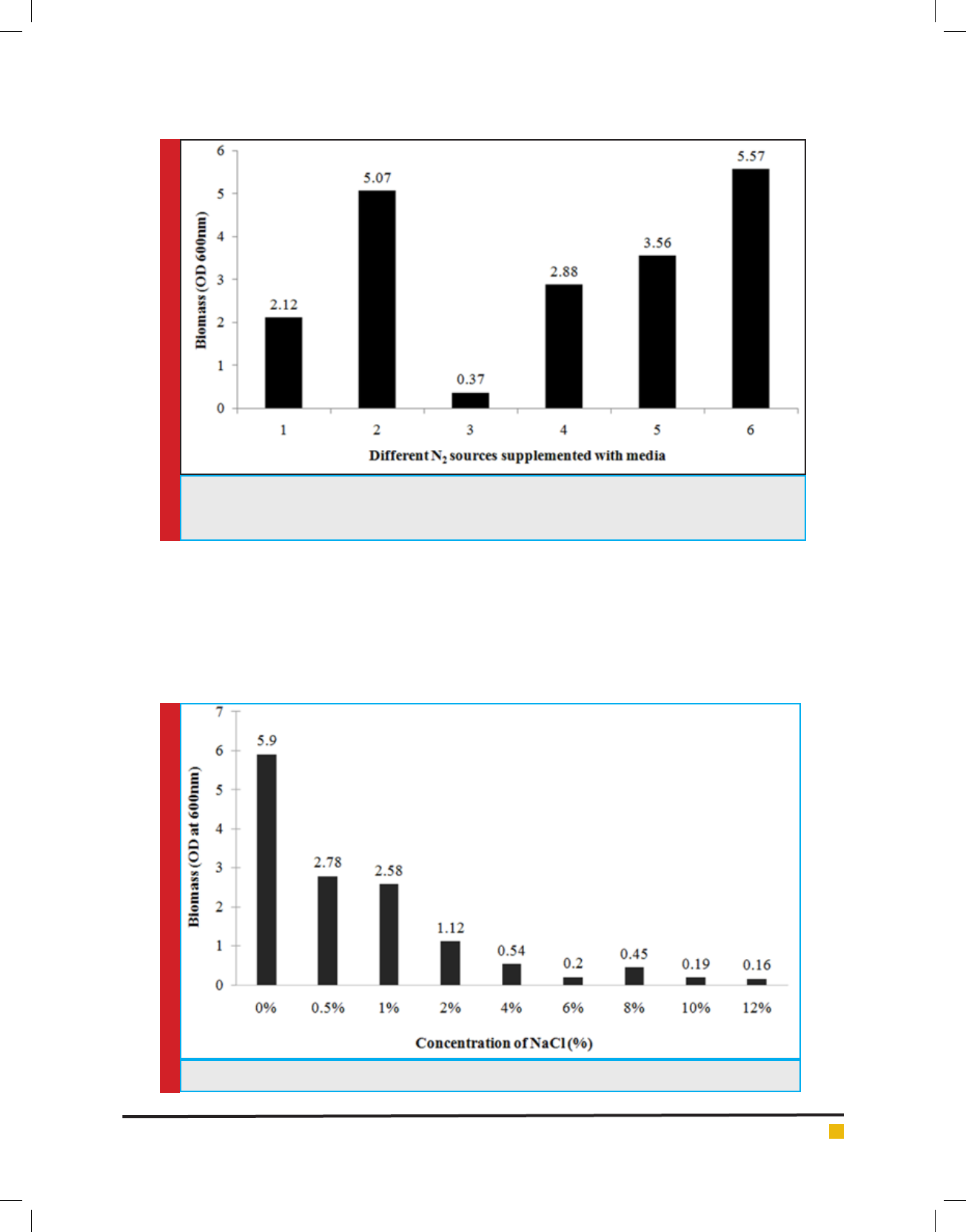
Effect of different nitrogen source on growth: Differ-
ent bacterial species utilize different nitrogen sources
for their growth. The different nitrogen sources selected
for this study are sodium nitrate, di-ammonium hydro-
gen citrate, ammonium nitrate, tryptophan type-1,
yeast extract, and ammonium chloride. TB media was
prepared to which different nitrogen source was added
separately at a nal concentration of 0.5%. The isolated
strain was inoculated in these media and grown under
37°C. The growth was measured after 24 hour of incuba-
tion. It was observed that Lysinibacillus sp. KEI-3 can
grow well in both organic as well as inorganic nitrogen
sources and the maximum biomass was formed in media
supplemented with yeast extract, tryptone and sodium
nitrite (Figure 5).
Effect of different salt concentration on growth: To
check the salt tolerance capacity, Lysinibacillus sp.KEI-3
was inoculated in TB medium supplemented with
various concentration of NaCl. The salt tolerance was
checked for concentration range of 0.5-12%. The strain
was inoculated and incubated at 37°C for 24 hours. After
incubation the biomass was measured spectrophotomet-
rically by reading OD at 600nm.Since isolates could not
grow well in Marine Broth (MB) due to its high salt con-
tent, therefore supplementing different concentrations
of NaCl in the medium was used to check maximum salt
tolerance of the Lysinibacillus sp.KEI-3which showed
tolerance till 1% and beyond that there was decline in
biomass formation. But it could grow till 12% with a
lower growth rate (Figure 6).
Submerged fermentation: Lysinibacillus sp.KEI-3 was
further subjected to submerged fermentation for pro-
duction of cellulase. Terri c Broth was supplemented
with optimized carbon and nitrogen sources (glucose/
sodium nitrite) that showed higher biomass for Lysini-
bacillus sp.KEI-3. Anaerobic fermentation was con-
tinued at 37°C for 4 days. Samples were collected and
FIGURE 4. Growth pro le of Lysinibacillus sp. KEI-3 strain with different carbon sources.
Shubhashree Mahalik et al.
BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS CELLULASE PRODUCTION IN
LYSINIBACILLUS
SP ISOLATED FROM THE ESTUARIES OF ODISHA 749
FIGURE 5. Growth pro le of Lysinibacillus sp. KEI-3 strain with different nitrogen sources. The numbers
on the horizontal axis represents various Nitrogen sources: 1.Ammonium chloride, 2.Sodium nitrite, 3.Di-
ammonium hydrogen citrate, 4.Ammonium nitrate, 5.Tryptone Type-I, 6.Yeast extract.
FIGURE 6. Growth pro le of Lysinibacillus sp. KEI-3 strain at different salt (NaCl) concentration.
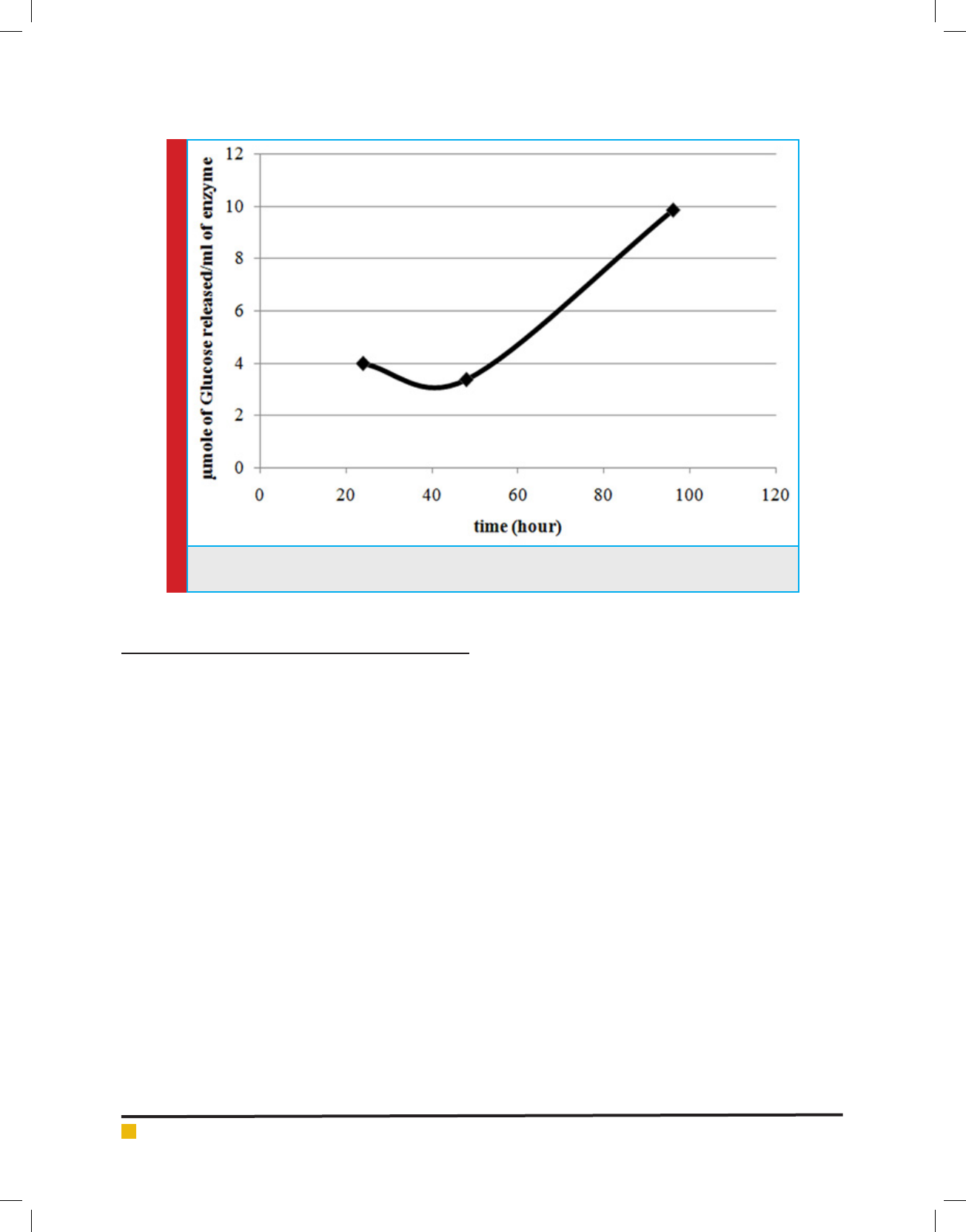
cellulase produced was quanti ed by DNS assay. The
results are represented in the form of µmole of glu-
cose (reducing equivalent) released per ml of cellulase
enzyme produced. Through the time course sampling, it
was observed that till 48 hours there was no signi cant
increase in cellulose production. Whereas, after 96 hours
of incubation there was 2.5 times increase in cellulose
production (Figure.7). This was a promising result and
the production could be increased by further scaling up
and optimization.
Shubhashree Mahalik et al.
750 CELLULASE PRODUCTION IN
LYSINIBACILLUS
SP ISOLATED FROM THE ESTUARIES OF ODISHA BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS
DISCUSSION
70% of the planet’s surface area is covered by oceans.
The coastal environments support huge diversity of
microbial life. But still only a small fraction of the spe-
cies has been cultured and identi ed till date due to cul-
ture related problems. Heavy pollution has led to severe
destruction of marine biological diversity (Abreo et al.,
2015; Baum et al., 2015).Trawler shing, pollution from
industries and drainage system has led to increase in
eutrophication which is further leading to change in
aquatic ecosystem as well as destruction of habitats.
This has led to decline in microbial biodiversity. So it
has become imperative to identify and isolate the organ-
isms and make a database, so that the information can
be evaluated time to time to check for the loss of species
in the marine world. Estuaries are dynamic in nature
in terms of the nutritional content and the associated
microbial population and this is mainly in uenced by
the convergence of fresh water and sea. The biochemical
environment of the estuaries makes it ideal for availabil-
ity of diverse microbial communities (Andersson et al.,
2014; Zhang et al., 2014). In different parts of the world
several studies have been undertaken to describe the
microbial diversity along the estuaries and their physi-
ochemical relation with the surrounding environment (
Reed & Martiny, 2013; Sun et al., 2014 and Lallias et al.,
2015).
Even though the eastern part of coastal India has sev-
eral estuaries, no substantial studies have been under-
taken to highlight the microbial diversity or the potenti-
alities for bioprospecting. In this work both the aspects
were covered, where pure cultures were isolated from
soil samples collected from estuaries and screened on
the basis of their ability to secrete cellulase enzyme.
Morphological and biochemical characterization of the
Isolate 3 showed similarity with Bacillus species. Phy-
logenetic analysis was done using 16s rRNA sequenc-
ing and the strain was identi ed to be Lysinibacillus
sp.KEI-3.These soil bacteria are rod shaped and gram
positive. Lysinibacillus fusiformis, Lysinibacillus spha-
ericus, Lysinibacillus boronitolerans, Lysinibacillus mac-
roides and Lysinibacillus xylanilyticus are some of the
strains that have been isolated and characterized previ-
ously. The strains belonging to this genus have several
industrial importance such as xylan degradation (Lee et
al., 2010), biodegradation of low-density polyethylene
(Esmaeili et al., 2013), biotransformation of Indole to
3-Methylindole (Arora et al., 2015), biological pest con-
trol (Rojas-Pinzón & Dussán, 2017).
These characters make this genus an interesting tar-
get for microbiological studies.The biomass production
FIGURE 7. Cellulase production pro le of Lysinibacillus sp. KEI-3 strain. Cellulase productivity is repre-
sented in the form of µmole of Glucose released per ml of cellulase produced.

BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS CELLULASE PRODUCTION IN LYSINIBACILLUS SP ISOLATED FROM THE ESTUARIES OF ODISHA 751
Shubhashree Mahalik et al.
was optimized under several physical and chemical
parameters such as temperature, media composition,
carbon source, nitrogen source and salinity tolerance.
The organisms showed high growth rate at 37ºC in TB
medium and SWGT medium. Bacterial species can uti-
lize both pentose and hexose sugars (Cook et al., 1993;
Kim et al., 2009; Liu et al., 2008). Each species has its
own ability to breakdown and utilize several carbon
sources. Also the preference of carbon source varies
from species to species (Brückner & Titgemeyer, 2002;
Görke & Stülke, 2008).Therefore the biomass yield was
assessed with various carbon sources which showed that
this species have a broad range of carbon preferences.
They could well utilize both pentose as well as hexose
sugars.
Organic as well as inorganic nitrogen sources are
critical for growth of microorganisms (Wheeler & Kirch-
man, 1986). The carbon to nitrogen (C/N) ratio is impor-
tant in a biological process (Cleveland & Liptzin, 2007).
Microorganisms require a proper nitrogen supplement
for metabolism during fermentation (Lin & Lay, 2004).
Therefore the optimal nitrogen was characterized in
order to obtain maximum biomass. It was observed that
Lysinibacillus sp.KEI-3 could produce higher biomass
when the media was supplemented with Sodium Nitrite.
Since the Isolates could not grow well in Marine Broth
(MB) due to its high salt content, therefore supplement-
ing different concentrations of NaCl in the medium was
used to check maximum salt tolerance. Lysinibacillus
sp.KEI-3 had higher salt tolerance and it can grow even
at 12 % NaCl.
Several Bacillus species are known to be cellulase
producers (Irfan et al., 2017; Sanjeev et al., 2017), but
very few reports are available on cellulase production
by Lysinibacillus strains (Khianngam et al., 2014). There-
fore submerged fermentation was done under the above-
optimized conditions and it was observed that the cell
could accumulate signi cant amounts of cellulase. Even
though the total units of cellulase produced are low as
compared to reported species, but it is a positive sign
that the isolated strain is a cellulase producer. Further
optimization of physical as well as bioprocess param-
eters could lead to accumulation of higher levels of cel-
lulase at high cell density cultures.
In the present scenario, the carbohydrolytic bacteria
or the lignocellulose degrading bacteria have a greater
industrial demand given their application in sacchari -
cation of lignocellulose for biofuel production. In this
context cellulase is the most common enzyme used in
the cocktail for degradation of lignocellulosic biomass
(Sindhu et al., 2016). Therefore there has been a continu-
ous effort to screen and isolate ef cient cellulase pro-
ducing microbes. These strains are isolated from various
geographical areas and also grown under various cheap
and alternative substrates for production of enzymes.
Apart from characterization of bacteria, process param-
eters as well as culture conditions are also being opti-
mized to enhance bacterial biomass to increase the
yield of cellulase. Optimization of media, carbon, nitro-
gen, salinity as well as temperature requirement led to
increased production of cellulase in the Lysinibacillus
sp.KEI-3.This isolated strain could further be screened
for other enzymes like xylanase and pectinase, which
would make it a potential strain for sacchari cation of
lignocellulose biomass.
ACKNOWLEDGMENT
The authors acknowledge the Bioprocess and Biosystems
Engineering lab, JNU, New Delhi for providing all neces-
sary help. Also P.G. Department of Biosciences and Bio-
technology, Fakir Mohan University, Balasore, Odisha is
acknowledged for providing infrastructure for carrying
out the experiments.
DECLARATION OF INTEREST STATEMENT: The authors
declare that they have no competing interests.
DATA AVAILABILITY STATEMENT: GenBank accession
number for the 16s rRNA nucleotide sequence of Lysini-
bacillus sp. KEI-3 is MG029268.
REFERENCES
Abreo, N. A. S., Macusi, E. D., Cuenca, G. C., Ranara, C. T. B.,
Cardona, L., & Arabejo, G. (2015). Nutrient enrichment, sedi-
mentation, heavy metals and plastic pollution in the marine
environment and its implications on Philippine marine biodi-
versity: A Review. IAMURE International Journal of Ecology
and Conservation, 15(1), 111-167.
Andersson, M. G. I., Berga, M., Lindström, E. S., & Langen-
heder, S. (2014). The spatial structure of bacterial communities
is in uenced by historical environmental conditions. Ecology,
95(5), 1134-1140.
Arora, P. K., Dhar, K., Garc, V., #xed, a, R. A., & Sharma, A.
(2015). Biotransformation of Indole to 3-Methylindole by Lysin-
ibacillus xylanilyticus Strain MA. Journal of Chemistry, 2015.
Baum, G., Januar, H. I., Ferse, S. C. A., & Kunzmann, A. (2015).
Local and Regional Impacts of Pollution on Coral Reefs along
the Thousand Islands North of the Megacity Jakarta, Indonesia.
PLOS ONE, 10(9), e0138271.
Bomble, Y. J., Lin, C.-Y., Amore, A., Wei, H., Holwerda, E.
K., Ciesielski, P. N., . . . Himmel, M. E. (2017). Lignocellulose
deconstruction in the biosphere. Current Opinion in Chemical
Biology, 41(Supplement C), 61-70.
Brückner, R., & Titgemeyer, F. (2002). Carbon catabolite repres-
sion in bacteria: choice of the carbon source and autoregula-
tory limitation of sugar utilization. FEMS microbiology letters,
209(2), 141-148.

752 CELLULASE PRODUCTION IN LYSINIBACILLUS SP ISOLATED FROM THE ESTUARIES OF ODISHA BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS
Shubhashree Mahalik et al.
Campbell, B. J., & Kirchman, D. L. (2013). Bacterial diversity,
community structure and potential growth rates along an estu-
arine salinity gradient. The ISME Journal, 7(1), 210-220.
Chun, J., Lee, J.-H., Jung, Y., Kim, M., Kim, S., Kim, B. K., &
Lim, Y.-W. (2007). EzTaxon: a web-based tool for the iden-
ti cation of prokaryotes based on 16S ribosomal RNA gene
sequences. International journal of systematic and evolution-
ary microbiology, 57(10), 2259-2261.
Cleveland, C. C., & Liptzin, D. (2007). C:N:P stoichiometry in
soil: is there a “Red eld ratio” for the microbial biomass? Bio-
geochemistry, 85(3), 235-252. doi:10.1007/s10533-007-9132-0
Coker, J. A. (2016). Extremophiles and biotechnology: current
uses and prospects. F1000Research, 5, F1000 Faculty Rev-
1396. doi:10.12688/f1000research.7432.1
Cook, G. M., Janssen, P. H., & Morgan, H. W. (1993). Simul-
taneous uptake and utilisation of glucose and xylose by
Clostridium thermohydrosulfuricum. FEMS microbiology let-
ters, 109(1), 55-61.
Esmaeili, A., Pourbabaee, A. A., Alikhani, H. A., Shabani, F.,
& Esmaeili, E. (2013). Biodegradation of Low-Density Polyeth-
ylene (LDPE) by Mixed Culture of Lysinibacillus xylanilyticus
and Aspergillus niger in Soil. PLOS ONE, 8(9), e71720.
Frank, J. A., Reich, C. I., Sharma, S., Weisbaum, J. S., Wilson,
B. A., & Olsen, G. J. (2008). Critical Evaluation of Two Prim-
ers Commonly Used for Ampli cation of Bacterial 16S rRNA
Genes. Applied and Environmental Microbiology, 74(8), 2461-
2470.
Görke, B., & Stülke, J. (2008). Carbon catabolite repression in
bacteria: many ways to make the most out of nutrients. Nature
Reviews Microbiology, 6, 613.
Irfan, M., Mushtaq, Q., Tabssum, F., Shakir, H. A., & Qazi, J. I.
(2017). Carboxymethyl cellulase production optimization from
newly isolated thermophilic Bacillus subtilis K-18 for sacchari-
cation using response surface methodology. AMB Express,
7(1), 29.
Juturu, V., & Wu, J. C. (2014). Microbial cellulases: Engineer-
ing, production and applications. Renewable and Sustainable
Energy Reviews, 33(Supplement C), 188-203.
Karakasidou, K., Nikolouli, K., Amoutzias, G. D., Pournou, A.,
Manassis, C., Tsiamis, G., & Mossialos, D. (2018). Microbial
diversity in biodeteriorated Greek historical documents dating
back to the 19th and 20th century: A case study. Microbiolo-
gyOpen, e00596.
Khianngam, S., Pootaeng-on, Y., Techakriengkrai, T., & Tana-
supawat, S. (2014). Screening and identi cation of cellulase
producing bacteria isolated from oil palm meal. Journal of
Applied Pharmaceutical Science,4(4), 90.
Kim, J.-H., Shoemaker, S. P., & Mills, D. A. (2009). Relaxed
control of sugar utilization in Lactobacillus brevis. Microbiol-
ogy, 155(4), 1351-1359.
Kuhad, R. C., Deswal, D., Sharma, S., Bhattacharya, A., Jain, K.
K., Kaur, A., . . . Karp, M. (2016). Revisiting cellulase produc-
tion and rede ning current strategies based on major chal-
lenges. Renewable and Sustainable Energy Reviews, 55, 249-
272.
Kuhad, R. C., Gupta, R., & Singh, A. (2011). Microbial Cellulases
and Their Industrial Applications. Enzyme Research, 2011, 10.
Kumar, S., Sharma, N., & Pathania, S. (2017). Cost effective
production of cellulase using wheat bran from Bacillus subtilis
BM1 and encoding endo-beta-1, 4-glucanase producing gene.
Molecular biology and evolution, Res. Environ. Life Sci.10(6)
507-512
Kumar, S., Stecher, G., & Tamura, K. (2016). MEGA7: molecular
evolutionary genetics analysis version 7.0 for bigger datasets.
Molecular Biology and Evolution, 33(7), 1870-1874.
L Bergquist, P., W Morgan, H., & Saul, D. (2014). Selected
enzymes from extreme thermophiles with applications in bio-
technology. Current Biotechnology, 3(1), 45-59.
Lallias, D., Hiddink, J. G., Fonseca, V. G., Gaspar, J. M., Sung,
W., Neill, S. P., . . . Creer, S. (2015). Environmental metabar-
coding reveals heterogeneous drivers of microbial eukaryote
diversity in contrasting estuarine ecosystems. The ISME Jour-
nal, 9(5), 1208-1221.
Lee, C. S., Jung, Y.-T., Park, S., Oh, T.-K., & Yoon, J.-H. (2010).
Lysinibacillus xylanilyticus sp. nov., a xylan-degrading bacte-
rium isolated from forest humus. International journal of sys-
tematic and evolutionary microbiology, 60(2), 281-286.
Lin, C. Y., & Lay, C. H. (2004). Carbon/nitrogen-ratio effect on
fermentative hydrogen production by mixed micro ora. Inter-
national Journal of Hydrogen Energy, 29(1), 41-45.
Littlechild, J. A. (2015). Enzymes from Extreme Environments
and Their Industrial Applications. Frontiers in Bioengineering
and Biotechnology, 3, 161.
Liu, S., Skinner-Nemec, K. A., & Leathers, T. D. (2008). Lac-
tobacillus buchneri strain NRRL B-30929 converts a concen-
trated mixture of xylose and glucose into ethanol and other
products. Journal of industrial microbiology & biotechnology,
35(2), 75-81.
Meddeb-Mouelhi, F., Moisan, J. K., & Beauregard, M. (2014). A
comparison of plate assay methods for detecting extracellular
cellulase and xylanase activity. Enzyme and Microbial Tech-
nology, 66(Supplement C), 16-19.
Moyle, P. B., Lund, J. R., Bennett, W. A., & Fleenor, W. E. (2010).
Habitat variability and complexity in the upper San Fran-
cisco Estuary. San Francisco Estuary and Watershed Science,
8(3).
Poli, A., Finore, I., Romano, I., Gioiello, A., Lama, L., & Nico-
laus, B. (2017). Microbial Diversity in Extreme Marine Habitats
and Their Biomolecules. Microorganisms, 5(2).
Reed, H. E., & Martiny, J. B. H. (2013). Microbial composition
affects the functioning of estuarine sediments. The ISME Jour-
nal, 7(4), 868-879.
Rojas-Pinzón, P. A., & Dussán, J. (2017). Ef cacy of the veg-
etative cells of Lysinibacillus sphaericus for biological con-
trol of insecticide-resistant Aedes aegypti. Parasites & Vectors,
10(1), 231.

BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS CELLULASE PRODUCTION IN LYSINIBACILLUS SP ISOLATED FROM THE ESTUARIES OF ODISHA 753
Shubhashree Mahalik et al.
Sindhu, R., Binod, P., & Pandey, A. (2016). Biological pretreat-
ment of lignocellulosic biomass–An overview. Bioresource
technology, 199, 76-82.
Stierle, A. A., & Stierle, D. B. (2014). Bioactive Secondary
Metabolites from Acid Mine Waste Extremophiles. Natural
product communications, 9(7), 1037-1044.
Subudhi, H., & Patra, H. K. (2013). Mangrove forests of river
estuaries of Odisha, India. International Journal of Biodiversity
and Conservation, 5(8), 446-454.
Sujana, K. A., Saravanan, R., & Pandey, A. D. (2015). Distribu-
tion of Aquatic Macrophytes in Balasore District, Odisha. In M.
Rawat, S. Dookia, & C. Sivaperuman (Eds.), Aquatic Ecosystem:
Biodiversity, Ecology and Conservation (pp. 1-12). New Delhi:
Springer India.
Sun, Z., Li, G., Wang, C., Jing, Y., Zhu, Y., Zhang, S., & Liu, Y.
(2014). Community dynamics of prokaryotic and eukaryotic
microbes in an estuary reservoir. Scienti c Reports, 4, 6966.
Wheeler, P. A., & Kirchman, D. L. (1986). Utilization of inor-
ganic and organic nitrogen by bacteria in marine systems.
Limnology and Oceanography, 31(5), 998-1009.
Yin, J., Chen, J.-C., Wu, Q., & Chen, G.-Q. (2015). Halophiles,
coming stars for industrial biotechnology. Biotechnology
advances, 33(7), 1433-1442.
Zhang, W., Bougouffa, S., Wang, Y., Lee, O. O., Yang, J., Chan,
C., . . . Qian, P.-Y. (2014). Toward Understanding the Dynamics
of Microbial Communities in an Estuarine System. PLOS ONE,
9(4), e94449.