Biotechnological
Communication
Biosci. Biotech. Res. Comm. 11(3): 461-468 (2018)
Insights into the antagonism of
Lactobacillus
fermentum
curd isolate against Gram-positive and
Gram-negative pathogenic bacteria
Debashis Halder and Shyamapada Mandal*
Department of Zoology, University of Gour Banga, Malda-732103, India
ABSTRACT
This communication characterizes the curd isolate of lactic acid bacteria having the capacity to antagonize human
pathogenic bacteria. The commercially available curd, in sealed form in a plastic cup, was procured from Malda town
market (West Bengal state, India) and processed microbiologically, using de Man Rogosa Sharpe medium, for the
isolation of lactic acid bacteria. The pure bacteria culture obtained was identi ed, by phenotypic characterization
through conventional methods, as Lactobacillus fermentum, and designated as LMEM 22. The Lactobacillus fermen-
tum LMEM 22 curd isolate had mixed antibiotic susceptibility patterns, showing resistance (ZDI: ≤15 mm) to ami-
kacin, cipro oxacin, kanamycin, methicillin and vancomycin, sensitivity (ZDI: ≥ 21 mm) to ampicillin, amoxyclav,
gentamycin, cefotaxime, imipenem, meropenem and tetracycline, and intermediate susceptibility (ZDI: 16 – 20 mm)
to cfoxitin and trimethoprim. The L. fermentum LMEM22 antagonizes both gram-negative: Acinetobacter bauman-
nii, Pseudomonas aeruginosa, Escherichia coli, Proteus vulgaris, Klebsiella pneumoniae, Salmonella enterica serovar
Typhi, and gram-positive: Staphylococcus aureus, Bacillus cereus, Enterococcus faecalis, Listeria monocytogenes,
bacterial pathogens, following agar overlay (ZDI: 17±1.73 mm to 20±1.00 mm, for gram-positive, and 18±2.00 mm
to 33±2.65 mm for gram-negative bacteria) as well as agar-well diffusion (ZDI: 10.67±2.08 mm to 12±1.00 mm, for
gram-positive, and 13.00±2.65 mm to 18.00±3.00 mm, for gram-negative bacteria) techniques. The overall bacteri-
ocin activity (AU/ml) of Lactobacillus fermentum LMEM 22 for the test bacterial pathogens ranged 142.27 – 240.00,
and the ‘R’ value ranged 5.5 – 13.5. This study underlines the usefulness of locally available lactic acid bacteria in
designing the probiotic microorganisms for biotherapy.
KEY WORDS: LACTIC ACID BACTERIA,
LACTOBACILLUS FERMENTUM
, ANTAGONISTIC ACTIVITY, BACTERIOCIN ACTIVITY, PATHOGENIC
BACTERIA
461
ARTICLE INFORMATION:
*Corresponding Author: samtropmed@gmail.com
Received 18
th
Aug, 2018
Accepted after revision 29
th
Sep, 2018
BBRC Print ISSN: 0974-6455
Online ISSN: 2321-4007 CODEN: USA BBRCBA
Thomson Reuters ISI ESC / Clarivate Analytics USA and
Crossref Indexed Journal
NAAS Journal Score 2018: 4.31 SJIF 2017: 4.196
© A Society of Science and Nature Publication, Bhopal India
2018. All rights reserved.
Online Contents Available at: http//www.bbrc.in/
DOI: 10.21786/bbrc/11.3/15

462 INSIGHTS INTO THE ANTAGONISM OF
LACTOBACILLUS FERMENTUM
CURD ISOLATE BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS
Debashis Halder and Shyamapada Mandal
INTRODUCTION
Among the lactic acid bacteria (LAB), Lactobacillus spp.
are characteristically known as probiotics, meaning, as
per the de nition of FAO/WHO (2001), the ‘live microor-
ganisms which when administered in adequate amounts
confer a health bene t on the host’; LAB isolates are
bene cial in many ways, but, essentially by restrict-
ing the toxigenic bacterial growth in the gut (Podolsky,
1998). The FAO/WHO (2007) suggested that the probiotic
microorganisms must possess the capacity to display
the antagonistic activity against bacterial pathogens.
Among the large number of lactobacilli isolated from
various fermented foods, 42 isolates showed activity
against Escherichia coli, while 15 isolates had antibac-
terial activity against Klebsiella pneumoniae (Shehata
etal., 2016).
Nivien et al. (2016) isolated LAB from fermented
milk, identi ed the isolates by phenotypic characteri-
zation and reported their antibacterial activity against
the bacterial strains: Escherichia coli, Salmonella Typh-
imurium, Staphylococcus aureus and Listeria monocy-
togenes. It has been reported that the lactobacilli, includ-
ing Lactobacillus fermentum, had growth inhibitory
action against gram-positive as well as gram-negative
human pathogenic bacteria (Vuotto etal., 2016). As per
the report of Sharma etal. (2016), the LAB (Pediococcus
acidilactici and Lactobacillus casei) isolated from milk
cream and lassi had growth inhibitory activity against a
number of gram-positive food borne bacteria.
Benavides etal. (2016) demonstrated that the Lacto-
bacillus fermentum isolate from local ecological niche
was sensitive to ampicillin, cefuroxime, tetracycline and
amoxicillin/clavulanic acid and resistant to gentamycin
and kanamycin, and the LAB was found inhibitory to
Escherichia coli and Salmonella Typhimurium. As per
the previous report (Halder et al., 2017), four lactoba-
cilli (Lactobacillus animalis LMEM6, Lactobacillus plan-
tarum LMEM7, Lactobacillus acidophilus LMEM8 and
Lactobacillus rhamnosus LMEM9) procured from differ-
ent commercially available curd samples had antibacte-
rial activity against gram-negative pathogenic bacteria,
such as Escherichia coli, Proteus vulgaris, Acinetobacter
baumannii and Salmonella enterica serovar Typhi, and
had resistance to vancomycin and amoxyclav.
Recently, Mahalot and Mandal (2018) have isolated
LAB from locally available cow milk and goat milk sam-
ples showing sensitivity to most of the test antibiotics,
while resistance was recorded for all isolates to methicil-
lin, forLactobacillussp. G1 andLactococcus sp. G2 to
trimethoprim, while to vancomycin forLactobacillussp.
G1 andLactobacillussp. C1. Since the good LAB are not
even waived from antibiotic resistance phenomenon,
many authors documented the status of various resist-
ances to antibiotics for safety pro ling of native LAB
isolates (Mandal etal., 2017). This background prompted
us to assess the broad spectrum antibacterial activity of
lactic acid bacillus isolated from commercially available
curd, and explore the antibiogram of the isolated Lacto-
bacillus, through phenotypic characterization.
MATERIAL AND METHODS
A single cup of commercially available curd sample
was procured from Malda town market (West Bengal,
India), and processed microbiologically for the isola-
tion of lactic acid bacteria (LAB), following the protocol
mentioned earlier (Halder and Mandal, 2015): growth
enrichment of LAB in MRS broth (Hi-Media, India), pure
culture (single discrete colony isolation) of LAB on MRS
agar (Hi-Media, India) plate, and storage of the LAB in
MRS agar stab at 4
o
C for further processing. The isolated
LAB (n=1), following Bergey’s manual (Holt, 1984), as
described earlier (Halder and Mandal, 2015), was sub-
jected to phenotypic (gram-staining, colony morphology
study and motility test) and biochemical (oxidase and
catalase production) characterization. The non-motile
non-spore forming gram-positive rod shaped bacteria
(no cocci were found), showing negative results to oxi-
dase and catalase tests, were subjected to IMViC, amino
acid decarboxylation and sugar fermentation tests.
The antibiotic susceptibility of the LAB (LMEM 22)
was executed by disc diffusion method (Bauer et al.,
1996), the details of which was described before (Hal-
der and Mandal, 2016; Halder etal., 2017). The antibi-
otic discs (Hi-Media, Mumbai, India) used in the study
included amikacin (Ak: 30-μg/disc), amoxyclav (Ac:
30-μg/disc), ampicillin (Am: 10-μg/disc), cipro oxacin
(Cp: 5-μg/disc), cfoxitin (Cx: 30-μg/disc), cefotaxime
(Ct: 30-μg/disc), cefotaxime/clavunilic acid (Cc: 30/10-
μg/disc), gentamycin (Gm: 30-μg/disc), imipenem (Ip:
10-μg/disc), kanamycin (Km: 30-μg/disc), methicil-
lin (Me: 5-μg/disc), meropenem (Mp: 10-μg/disc), tet-
racycline (Tc: 30-μg/disc), trimethoprim (Tm: 5-μg/
disc) and vancomycin (Vm: 30-μg/disc). The results, in
terms of ZDI (zone diameter of inhibition) values, were
interpreted according to Liasi etal. (2009) and Vlkova
etal. (2006), in order to label the test bacterial isolate
as resistant (ZDI: ≤ 15 mm), sensitive (ZDI: 21 mm), or
intermediately susceptible (ZDI: 16–20 mm).
The antagonistic activity of the LAB LMEM 22 isolate
from curd was determined against gram-negative (Aci-
netobacter baumannii, Pseudomonas aeruginosa, Escher-
ichia coli, Proteus vulgaris, Klebsiella pneumoniae, and
Salmonella enterica serovar Typhi) and gram-positive
(Staphylococcus aureus, Bacillus cereus, Enterococcus
faecalis, Listeria monocytogenes) bacterial pathogens,
BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS INSIGHTS INTO THE ANTAGONISM OF
LACTOBACILLUS FERMENTUM
CURD ISOLATE 463
Debashis Halder and Shyamapada Mandal
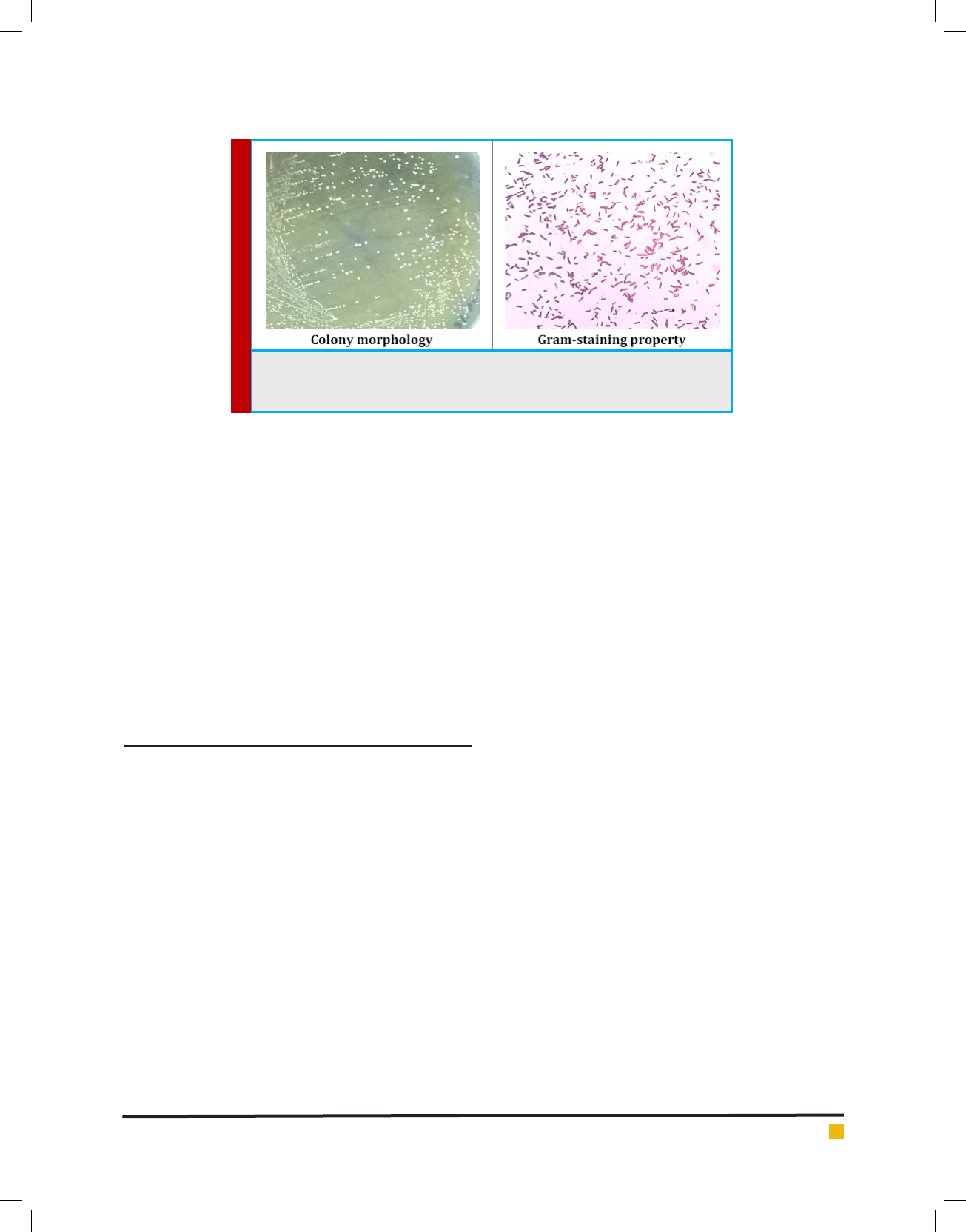
FIGURE 1. Colony morphology and gram-staining property of the isolated LAB
from curd sample. The LAB forms small white circular colonies on MRS agar plate
and the isolate is gram-positive rod, forming no spore.
following agar-well diffusion (Tagg, 1971; Halder etal.,
2017) and agar overlay (Shokryazdan etal., 2018) meth-
ods, as described and interpreted earlier (Shokryazdan
et al., 2014, Halder et al., 2017, Mandal and Halder,
2018).
The ‘R’ values, from the action of LMEM 22 isolate
over the bacterial pathogens, were calculated applying
the formula described elsewhere (Halder and Mandal,
2016), and interpreted according to the criteria men-
tioned earlier (Carasi etal., 2014; Pisano etal., 2014),
while the bacteriocin activity of LAB (LMEM 22), in
terms of arbitrary units per milliliter (AU/ml), was calcu-
lated following the formula put forwarded by Iyapparaj
etal. (2013).
RESULTS AND DISCUSSION
As has been demonstrated by Iyapparaj etal. (2013), the
morphologically identical bacterial colonies, procured
from goat milk on the MRS agar plate, have been iden-
ti ed as Lactobacillus sp., on the basis of physical and
biochemical characteristics, following Holt etal. (1984).
In this study, a single isolate of non-motile non-spore
forming gram-positive rod was procured from the curd
sample (Figure 1), and the isolated bacteria (LMEM 22)
was an hetero-fermentative strain, which in TSI test
showed the production of acid as well as gas (CO
2
). The
LMEM 22 isolate showed negative test results for cata-
lase and oxidase, and in IMViC test battery the isolates
was positive for methyl red. The sugar fermentation pat-
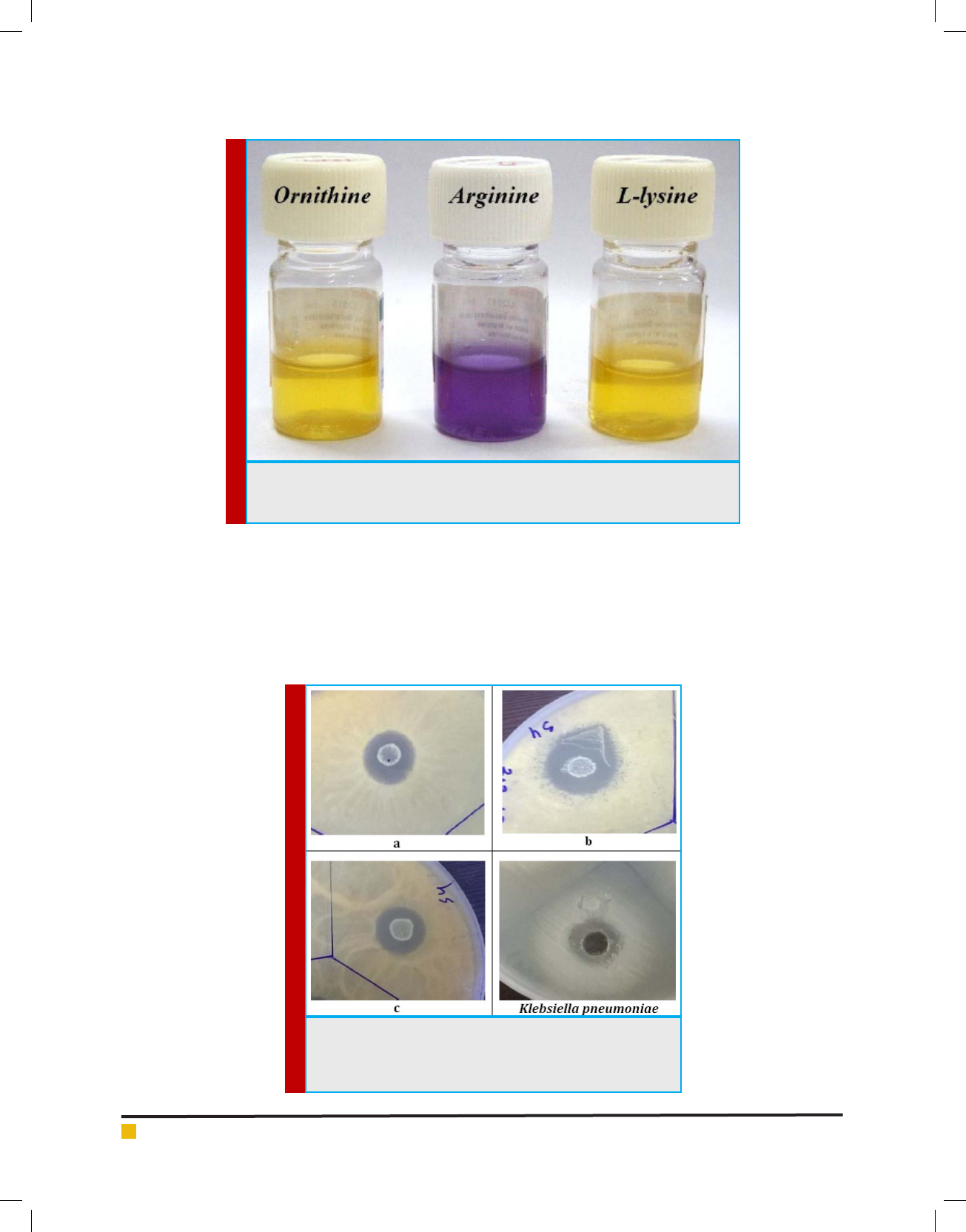
tern of LMEM 22 isolate is represented in Table 1, while
the amino acid decarboxylation test results are depicted
in Figure 2. Thus, following phenotypic and biochemi-
cal characterization the isolated LAB was identi ed as
Lactobacillus fermentum LMEM 22. The LAB isolates
procured from different fermented foods, including
curd, have been identi ed earlier by Nigam etal. (2012),
following phenotypic characterization of the bacteria.
Currently, the treatment options with antibiotics are
inadequate because of the escalating rate of emergence
of antibiotic resistant pathogenic bacteria causing life-
threatening infections to humans. Alternative to the
antibiotics, which remains the mainstay of all therapy
for bacterial infections (Van Boeckel etal., 2014), probi-
otic lactobacilli have been found suitable for biotherapy
with proven antibacterial activity (Iyapparaj etal., 2013;
Sing etal., 2017).
Earlier, it has been reported that the curd isolates of
Lactobacillus animalis LMEM6, Lactobacillus plantarum
LMEM7, Lactobacillus acidophilus LMEM8 and Lactoba-
cillus rhamnosus LMEM9, had bacterial growth inhibi-
tory activity, having ZDIs 13.67 ± 0.58 – 29.50 ± 2.10
mm, by agar-well, and 11.33 ± 0.58 – 35.67 ± 2.52, by
agar overlay, against human pathogenic bacteria, viz.,
Escherichia coli, Proteus vulgaris, Acinetobacter bau-
mannii and Salmonella enterica serovar Typhi (Halder
etal., 2017). The antibacterial activity of Lactobacillus
fermentum LMEM 22 against gram-positive and gram-
negative pathogenic bacteria, following agar-well diffu-
sion method is depicted in Figure 3.
Gandevia etal. (2017) isolated, from cow milk, buf-
falo milk, goat milk and curd samples, a number of Lac-
tobacillus species, including Lactobacillus fermentum,
having the capacity to inhibit the growth of gram-posi-
tive bacteria, such as Staphylococcus aureus (ZDI: 8 – 17
mm) and Bacillus cereus (ZDI: 12 – 22 mm). The two
Lactobacillus fermentum isolates from buffalo milk had
ZDI of 19 mm, while the Lactobacillus oris conferred
ZDI of 18 mm, against Listeria monocytogenes (Melia
etal., 2017). The broad spectrum antibacterial activity of
Lactobacillus fermentum has been demonstrated earlier
(Ilayajara etal., 2011; Ramasamy and Suyambulingam,
2015; Podolsky, 1998), in which the LAB showed growth
Debashis Halder and Shyamapada Mandal
464 INSIGHTS INTO THE ANTAGONISM OF
LACTOBACILLUS FERMENTUM
CURD ISOLATE BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS
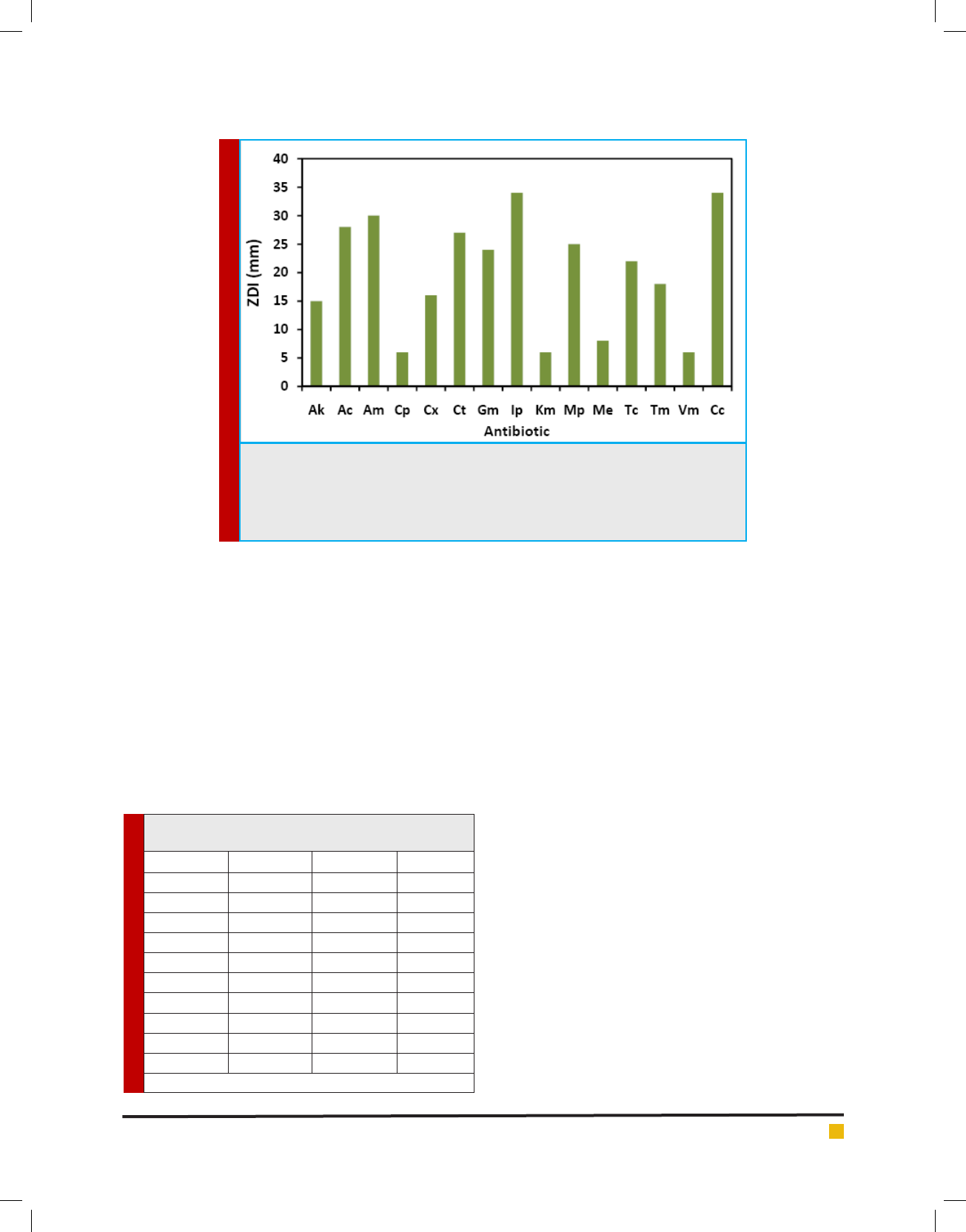
FIGURE 3. Antibacterial activity of Lactobacillusfermentum
LMEM22 isolate from curd sample; by Agar-overlay: (a) S. enter-
ica Typhi, (b) S. aureus, (c) Klebsiella pneumoniae; Agar-well
diffusion: (d) Klebsiella pneumoniae.
FIGURE 2. Amino acid decarboxylation test results for the isolated LAB from curd
sample. The LAB utilized arginine, but not ornithine and L-lysine, in Moeller decar-
boxylase broth (Hi-Media, India).
inhibitory activity against gram-positive (Staphylococ-
cus aureus, Enterococcus spp., Streptococcus spp., Bacil-
lus subtilis) and gram-negative (Proteus spp., Pseu-
domonas aeruginosa, Escherichia coli and Klebsiella
pneumoniae) bacterial pathogens.
Kang etal. (2017) reported the enhancing killing of
Staphylococcus aureus strains by Lactobacillus salivar-
ius except Staphylococcus aureus ATCC 25923 strain,
the growth of which was fully killed by Lactobacillus
fermentum.
Sharma et al. (2016) isolated two lactic acid bac-
teria: Pediococcus acidilactici and Lactobacillus casei
from milk cream and lassi, respectively, which had anti-
bacterial activity against a large number of potential
Debashis Halder and Shyamapada Mandal
BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS INSIGHTS INTO THE ANTAGONISM OF
LACTOBACILLUS FERMENTUM
CURD ISOLATE 465
FIGURE 4. Antibiotic susceptibility test results for Lactobacillus fermentum LMEM22
isolate. Ak: amikacin; Ac: amoxyclav; Am: ampicillin; Cp: cipro oxacin; Cx: cfoxi-
tin; Ct: cefotaxime; Cc: cefotaxime/clavunilic acid; Gm: gentamicin; Ip: imipenem; Km:
kanamycin; Me: methicillin; Mp: meropenem; Tc: tetracycline; Tm: trimethoprim; Vm:
vancomycin.
Table 1. Sugar fermentation test results for the isolated
LAB from curd sample
Sugars Utilization Sugars Utilization
Adonitol - D-Melezitose +
Arabinose W Raf nose +
Cellobiose + Rhamnose +
Dextrose + D-Ribose +
Esculin - Salicin -
Glucose + (g) Sorbitol +
Lactose + Sucrose +
Mannitol + Trehalose +
Mannose + Xylose +
Melibiose + D-Galactose +
+: Strong fermentation; w: week fermentation –: No fermentation
food-borne bacteria, viz., Staphylococcus aureus, Ente-
rococcus faecalis, Listeria monocytogenes, Clostridium
perfringens and Bacillus cereus having respective ZDIs
range from 12 mm to 20 mm and from 13 mm to 19 mm.
In the current study, the agar-well diffusion had ZDIs
of 10.67±2.08 mm (Staphylococcus aureus) to 12±1.00
mm (Bacillus cereus), for gram-positive, and 13.00±2.65
mm (Escherichia coli) to 18.00±3.00 mm (Salmonella
enterica Typhi), for gram-negative bacteria (Table 2).
The Lactobacillus fermentum LMEM 22 isolate, follow-
ing agar overlay method, also had antibacterial activity
against the indicator bacterial strains (Figure 3), display-
ing ZDIs from 17.00±1.73 mm (Listeria monocytogenes)
to 20.00±1.00 mm (Enterococcus faecalis), for gram-
positive, and from 18.00±2.00 mm (Escherichia coli) to
33±2.65 mm (Acinetobacter baumannii) for gram-nega-
tive bacteria (Table 3).
The bacteriocin activity of the isolated LAB, Lacto-
bacillus fermentum LMEM 22, has been shown in Table
2, while the Table 3 depicts the ‘R’ values of the iso-
lated LAB. Iyapparaj etal. (2013) isolated Lactobacillus
sp. MSU3IR strain, which against pathogenic bacteria:
Staphylococcus aureus and Pseudomonas aeruginosa,
displayed higher bacteriocin activity (393.2 ± 2.61 to
556.0 ± 5.34 AU/ml) in MRS medium, compared to the
activity value (341.2 ± 2.36 to 473.2 ± 3.96 AU/ml) as
recorded in Lactobacillus selection broth. Earlier, the ‘R’
values of curd lactobacilli strains ranged 3.00 – 13.17
mm, while the bacteriocin activity, in terms of arbitrary
units (AU/ml), ranged 155.60 – 293.33, against MDR
Klebsiella pneumoniae clinical isolates (Mandal and Hal-
der, 2018).
In another study, the probiotic lactobacilli had excel-
lent antibacterial activity against gram-negative human
pathogenic bacteria (Escherichia coli, Proteus vulgaris,
Acinetobacter baumannii and Salmonella enterica sero-
var Typhi) displaying ‘R’ values ranging from 3.17 ±
0.29 to 15.33 ± 1.26 mm, and the bacteriocin activity
ranging from 233.34 ± 45.54 to 280.56 ± 83.67 AU/ml
(Halder etal., 2017). As per the report of Shehata etal.
(2016), among nine isolates of LAB, one (Lactococcus
lactic subsp. lactis) had strong activity (1600 AU/ml)

Debashis Halder and Shyamapada Mandal
466 INSIGHTS INTO THE ANTAGONISM OF
LACTOBACILLUS FERMENTUM
CURD ISOLATE BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS
Table 2. Agar-well diffusion test results in terms ZDI (mm) and the calculated bacteriocin
activity (Au/ml) of Lactobacillus fermentum LMEM22 against gram-positive and gram-
negative indicator bacterial strains.
Indicator strains Bacterial isolates ZDI range (Mean ± SD) Bacteriocin activity
Gram-negative E. coli 10-15 (13.00±2.65) 173.33
Pr. vulgaris 12-14 (13.33±1.15) 177.73
A. baumannii 16-20 (17.33±2.31) 231.07
Ps. aeruginosa 15-16 (15.67±0.58) 208.93
K. pneumoniae 14-17 (15.67±1.53) 208.93
S. enterica Typhi 15-21 (18.00±3.00) 240.00
Gram-positive B. cereus 11-13 (12.00±1.00) 160.00
E. faecalis 10-13 (11.67±1.53) 155.60
S. aureus 9-13 (10.67±2.08) 142.27
L. monocytogenes 10-13 (11.67±1.53) 155.60
Au/ml: arbitrary units per milliliter
Table 3. Agar overlay test results in terms ZDI (mm) and the calculated ‘R’ values for
Lactobacillus fermentum LMEM22 against gram-positive and gram-negative indicator
bacterial strains
Indicator strains Bacterial isolates ZDI range (Mean ± SD) ‘R’ value (mm)
Gram-negative E. coli 16-20 (18.00±2.00) 6.00
Pr. vulgaris 21-26 (23.33±2.52) 8.67
A. baumannii 31-36 (33.00±2.65) 13.50
Ps. aeruginosa 23-28 (26.00±2.65) 10.00
K. pneumoniae 20-25 (22.67±2.52) 8.34
S. enterica Typhi 20-25 (22.00±2.65) 8.00
Gram-positive B. cereus 16-19 (17.67±1.53) 5.84
E. faecalis 19-21 (20.00±1.00) 7.00
S. aureus 18-20 (18.67±1.56) 6.34
L. monocytogenes 15-18 (17.00±1.73) 5.50
SD: standard deviation; ZDI: zone diameter of inhibition.
against Klebsiella pneumoniae, while, four isolates had
bacteriocin activity of 800 AU/ml against Escherichia
coli (for Lactobacillus paracasei), Streptococcus pyogenes
(for Lactobacillus gasseri), Staphylococcus aureus (for
Lactobacillus rhamnosus) and Salmonella senftenberg
(for Lactobacillus gasseri RM28). The current investiga-
tion demonstrates the capacity of antibacterial activity
of Lactobacillus fermentum LMEM 22 for the indicator
microorganisms, consisting of both gram-positive and
gram-negative pathogenic bacteria, with an overall bac-
teriocin activity (AU/ml) of 142.27 – 240.00 and the ‘R’
values of 5.5 – 13.5 mm.
The antibiotic susceptibility test results for Lactoba-
cillus fermentum LMEM 22 isolate is shown in Figure
4. One of the most important probiotic features, de n-
ing safe for human consumption, of lactic acid bacte-
ria is being their antibiotic sensitivity, and the intrinsic
resistance (chromosomally conferred from point muta-
tion) property as well (Georgieva et al., 2015). As has
been demonstrated by Benavides etal. (2016), the iso-
lated Lactobacillus fermentum was sensitive to ampicil-
lin (ZDI: 28 mm), cefuroxime (ZDI: 30 mm), tetracycline
(ZDI: 24 mm) and amoxicillin/clavulanic acid (ZDI: 26
mm) and resistant to gentamycin and kanamycin, and
the LAB was found inhibitory to Escherichia coli (ZDI:
13 mm) and Salmonella Typhimurium (ZDI: 12 mm). As
per our earlier report the curd lactobacilli had sensitiv-
ity to majority of the test antibiotics displaying a com-
mon resistance to Vm (Halder and Mandal, 2016). In the
instant case, the isolated LAB: Lactobacillus fermentum
LMEM 22 showed resistance to Ak, Cp, Km, Me and Vm
(ZDI: ≤15 mm; range: 6 – 15 mm), and such resistances
are intrinsic as well as non-transferable (Bamidele etal.,
2017; Imperial and Ibana, 2016).
The Lactobacillus fermentum LMEM 22 was sensitive
to Ac, Am, Cc, Cx, Gm, Im, Mp and Tc (ZDI: 21 mm;

Debashis Halder and Shyamapada Mandal
BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS INSIGHTS INTO THE ANTAGONISM OF
LACTOBACILLUS FERMENTUM
CURD ISOLATE 467
range: 22 – 34 mm), and intermediately susceptible to
Cx and Tm with ZDIs of 16 and 18 mm (ZDI criteria
range: 16 – 20 mm); the LAB, while, showed resistance
(ZDI: ≤15 mm) to Ak, Cp, Km, Me and Vm. Thus, the
isolated LAB, in this study, has been found to be safe,
on the basis of lack of transferable antibiotic resistance
property (Ammor etal., 2008; Imperial and Ibana, 2016),
and this LAB might be useful in single-strain based pro-
biotic formulation bene ting a large number of local
population, in this part of the globe. It has been reported
that multi-strain/multi-general probiotics might exhibit
limited functional property for universal usage, requir-
ing probiotics alternatives development and/or person-
alized probiotic approaches (Zmora et al., 2018; Suez
etal., 2018).
CONCLUSION
The Lactobacillus fermentum LMEM22, which was iso-
lated from locally available commercial curd, showed
antagonistic activity against gram-positive as well as
gram-negative pathogenic bacteria, with overall bacte-
riocin activity (AU/ml) of 142.27 – 240.00, and the ‘R’
value of 5.5 – 13.5, and (based upon the report available
in literatures, too) there is no risk of transferable antibi-
otic resistance in the LAB. Thus, the isolated LAB might
be useful as broad spectrum antibacterial biotherapeu-
tics, and such native LAB isolate might be consumed
alone, in place of antibiotic therapy, or can be used
(based upon the antibiogram of the native LAB) in pro-
biotic-antibiotic combination therapy. However, further
studies are needed to validate the probiotic attributes of
the isolated LAB, including its molecular identity as well
as the antibiotic resistance management.
REFERENCES
Ammor, M.S., Florez, A.B., van Hoek, A.H., de los Reyes-
Gavilan, C.G., Aarts, H.J., Margolles, A and Mayo, B. (2008).
Molecular characterization of intrinsic and acquired antibiotic
resistance in lactic acid bacteria and bi dobacteria. Journal of
Molecular Microbiology and Biotechnology, 14:6–15.
Bamidele, T.A., Adeniyi, B.A and Fowora, M.J. (2017). Anti-
biotic resistance patterns of lactic acid bacteria isolated from
Nigerian grown salad vegetables. African Journal of Microbi-
ology Research, 11: 433-439.
Bauer, A.J., Kirby, W and Turck, M. (1996). Antibiotic suscep-
tibility testing by standardized single disc method. American
Journal of Clinical Pathology, 45: 493–496.
Benavides, A.B., Ulcuango, M., Yepez, L and Tenea, G.N.
(2016). Assessment of the in vitro bioactive properties of lactic
acid bacteria isolated from native ecological niches of Ecuador.
Revista Argentina De Microbiologia, 48:236-244.
Carasi, P., Diaz, M., Racedo, S.M., Antoni, G.D., Urdaci, M.C
and Serradell, M.A. (2014). Safety characterization and antimi-
crobial properties of ke r-isolated Lactobacillus ke ri. Biomed
Research International, 2: 1–7.
FAO/WHO (2001). Health and nutritional properties of pro-
biotics in food including powder milk with live lactic acid
bacteria: report of a Joint FAO/WHO expert consultation on
evaluation of health and nutritional properties of probiotics
in food including powder milk with live lactic acid bacteria,
Technical Report, Food and Agriculture Organization/World
Health Organization, Cordoba, Argentina. http://www.who.int/
foodsafety/publications/fs management/en/probiotics.pdf.
FAO and WHO, 2007. WHO working group on drafting guide-
lines for the evaluation of probiotics in food. Guidelines for the
evaluation of probiotics in food: Report of a joint FAO/WHO.
FAO and WHO, London, Ontario, Canada.
Gandevia, H., Rana, N and Desai, B. (2017). Screening, produc-
tion and antibacterial activity of bacteriocin from Lactobacil-
lus spp. BMR Microbiology, 3: 1-8.
Georgieva, R., Yocheva, L., Tserovska, L., Zhelezova, G., Ste-
fanova, N and Atanasova, A. (2015). Antimicrobial activity
and antibiotic susceptibility of Lactobacillus and Bi dobacte-
rium spp. intended for use as starter and probiotic cultures.
Biotechnology and Biotechnological Equipment, 29: 84–91.
Halder, D and Mandal, S. (2015). Curd lactobacilli with probi-
otic potentiality. Translational Biomedicine, 6:1-6.
Halder, D and Mandal, S. (2016). Antibacterial potentiality of
commercially available probiotic lactobacilli and curd lactoba-
cilli strains, alone and in combination, against human patho-
genic bacteria. Translational Biomedicine, 7: 1–7.
Halder, D., Mandal, M., Chatterjee, S.S., Pal, N.K and Mandal,
S. (2017). Indigenous probiotic lactobacillus isolates present-
ing antibiotic like activity against human pathogenic bacteria.
Biomedicines, 5: 1-11.
Mandal, S and Halder, D. (2018). Exploring Anti-Klebsiella
pneumoniae activity of probiotic lactobacilli of curd origin.
Acta Scienti c Microbiology, 1: 49-53.
Holt, J.G and Krieg, N.R. (1984). Bergey’s Manual of System-
atic Bacteriology; Williams and Wilkins: Baltimore, MD, USA.
Ilayajara, R., Radhamadhavan, P.A and Nirmala, C.B. (2011).
Assessment for potential use of bacteriocin producing Lac-
tobacillus fermentum isolated from human milk for prevent-
ing urinary tract infections. Journal of Pharmacy Research, 4:
3445–3447.
Imperial, I.C.V.J and Ibana, J.A. (2016). Addressing the antibi-
otic resistance problem with probiotics: reducing the risk of its
double-edged sword effect. Frontiers in Microbiology, 7: 1-10.
Iyapparaj, P., Maruthiah, T., Ramasubburayan, R., Prakash, S.,
Kumar, C., Immanuel, G and Palavesam, A. (2013). Optimiza-
tion of bacteriocin production by Lactobacillus sp. MSU3IR
against shrimp bacterial Pathogens. Aquatic Biosystems, 9:1-
10.
Kang, M.S., Lim, H.S., Jong-Suk Oh, J.K., Lim, Y.J., Wuertz-
Kozak, K., Harro, J.M., Mark E. Shirtliff, M.E and Achermann,

Debashis Halder and Shyamapada Mandal
468 INSIGHTS INTO THE ANTAGONISM OF
LACTOBACILLUS FERMENTUM
CURD ISOLATE BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS
Y. (2017). Antimicrobial activity of Lactobacillus salivarius
and Lactobacillus fermentum against Staphylococcus aureus.
Pathogens and Disease, 75: 1-10.
Liasi, S.A., Azmi, T.I., Hassan, M.D., Shuhaimi, M and Rosfari-
zan, M. (2009). Antimicrobial activity and antibiotic sensitiv-
ity of three isolates of lactic acid bacteria from fermented sh
product Budu. Malaysian Journal of Microbiology, 5: 33–37.
Mahalot, A and Mandal, S. (2018). Assessment of lactic acid
bacteria from cow milk and goat milk samples for probiotic
potentiality by in vitro methods. Acta Scienti c Pharmaceuti-
cal Sciences, 2: 56-58.
Melia, S., Purwati, E., Yuherman, Jaswandi, Aritonang, S.N and
Silaen, M. (2017). Characterization of the antimicrobial activ-
ity of lactic acid bacteria isolated from buffalo milk in west
Sumatera (Indonesia) againstListeria monocytogenes.Pakistan
Journal of Nutrition, 16: 645-650.
Nigam, A., Kumar, A., Madhusudan, H.V and Bhola, N. (2012).
In-vitro screening of antibacterial activity of lactic acid bacte-
ria against common enteric pathogens. Journal of Biomedical
Sciences, 1: 1-6.
Nivien, A., Ghani, S.A.E., Gomaa, R.S and Fouad, M.T. (2016).
Molecular identi cation of potential probiotic lactic acid bac-
teria strains isolated from Egyptian traditional fermented dairy
products. Biotechnology, 15: 35-43.
Pisano, M.B., Viale, S., Conti, S., Fadda, M., Deplano, M., Melis,
M.P., Deiana, M and Cosentino, S. (2014). Preliminary evalu-
ation of probiotic properties of Lactobacillus strains isolated
from Sardinian dairy products. Biomed Research International,
2: 1–8.
Podolsky, S. (1998). Cultural divergence: Elie Metchnikoff’s
Bacillus bulgaricus therapy and his underlying concept of
health. Bulletin of the History of Medicine, 72: 1–27.
Ramasamy, T.K and Suyambulingam, K. (2015). Molecular
characterization of Lactobacillus sp. from Indian curd and its
antagonistic effects on uropathogens of diabetic patients. Inter-
national Research Journal of Biological Sciences, 4: 12–22.
Sharma, K., Sharma, N and Sharma, R. (2016). Identi cation
and evaluation of in vitro probiotic attributes of novel and
potential strains of lactic acid bacteria isolated from traditional
dairy products of north-west Himalayas. Journal of Clinical
Microbiology and Biochemical Technology, 2: 018-025.
Shehata, M.G., Sohaimy S.A.E., Malak, A and Youssef, E.M.M.
(2016). Screening of isolated potential probiotic lactic acid
bacteria for cholesterol lowering property and bile salt hydro-
lase activity. Annals of Agricultural Science, 61: 65–75.
Shokryazdan, P., Sieo, C.C., Kalavathy, R., Liang, J.B., Alitheen,
N.B., Jahromi, M.F and Ahmed, M. (2014). Probiotic poten-
tial of Lactobacillus strains with antimicrobial activity against
some human pathogenic strains. BioMed Research Interna-
tional, 2: 1–16.
Singh, B., Mal, G and Marotta, F. (2017). Designer probiotics:
paving the way to living therapeutics. Trends in Biotechnol-
ogy, 35: 679-681.
Suez, J., Zmora, N., Zilberman-Schapira, G., Mor, U., Dori-
Bachash, M., Bashiardes, S., Zur, M., Regev-Lehavi, D., Brik,
R.B.-Z., Federici, S., etal. (2018). Post-antibiotic gut mucosal
microbiome reconstitution is impaired by probiotics and
improved by autologous FMT. Cell 174, 1406–1423.
Tagg, J.R and McGiven, A.R. (1971). Assay system for bacteri-
ocins. Applied Microbiology, 21: 943–944.
Van Boeckel, T.P., Gandra, S., Ashok, A., Caudron, Q., Grenfell,
B.T., Levin, S.A and Laxminarayan, R. (2014). Global antibiotic
consumption 2000 to 2010: an analysis of national pharma-
ceutical sales data. Lancet Infectious Disease, 14: 742–750.
Vlkova, E., Rada, V., Popelarova, P., Trojanová, I and Killer, J.
(2006). Antimicrobial susceptibility of bi dobacteria isolated
from gastrointestinal tract of calves. Livestock Science, 105:
253–259.
Vuotto, C., Longo, F and Donelli, G. (2014). Probiotics to coun-
teract bio lm-associated infections: promising and con icting
data. International Journal of Oral Science, 6: 189–194.
Zmora, N., Zilberman-Schapira, G., Suez, J., Mor, U., Dori-
Bachash, M., Bashiardes, S., Kotler, E., Zur, M., Regev-Lehavi,
D., Brik, R.B.-Z., etal. (2018). Personalized gut mucosal col-
onization resistance to empiric probiotics is associated with
unique host and microbiome features. Cell 174, 1388–1405.