
Medical
Communication
Biosci. Biotech. Res. Comm. 10(1): 236-240 (2017)
Association of gene variants with longitudinal
changes in high density lipoprotein cholesterol among
Tehranian people: A latent growth curve model
Mahdi Akbarzadeh
1
, Abbas Moghimbeigi
2
*, Nathan Morris
3
, Maryam S Daneshpour
4
,
Ali Reza Soltanian
5
and Hossein Mahjub
6
1
Ph.D Candidate in Biostatistics, Department of Biostatistics, School of Public Health, Hamadan University
of Medical Sciences, Hamadan, Iran
2
Associate Professor of Biostatistics, Modeling of Noncommunicable Disease Research Canter, Department of
Biostatistics, School of Public Health, Hamadan University of Medical Sciences, Hamadan, Iran
3
Associate Professor of Biostatistics, Department of Epidemiology and Biostatistics, Case Western Reserve
University, Cleveland, OH, USA
4
Cellular and Molecular Endocrine Research Center, Research Institute for Endocrine Sciences, Shahid
Beheshti University of Medical Sciences; Tehran, Iran
5
Associate Professor of Biostatistics, Modeling of Noncommunicable Disease Research Canter, Department of
Biostatistics, School of Public Health, Hamadan University of Medical Sciences, Hamadan, Iran
6
Professor of Biostatistics, Research Center for Health Sciences and Department of Biostatistics, School of
Public Health, Hamadan University of Medical Sciences, Hamadan, Iran
ABSTRACT
High density lipoprotein cholesterol (HDL-C) particles remove fat molecules from cells which need to export fat molecules. The aim
of this study was to assess the association of SNP related to fat mass and obesity associated gene (FTO) with HDL-C change in a
subset of participated families in Tehran Lipid and Glucose Study (TLGS). In this study 914 individual (with age>3) including 126
families with size 8.49±3.10 were selected from participants in TLGS. Genomic DNA was extracted from peripheral blood using
standard salting-out method. HDL-C measured in four time point from March 1999 to December 2011 with a 3-year follow-up
period. We examine the association via a latent growth curve model. To achieve the study target we used the latent growth curve
model (LGCM) in R in TLGS family data. Adjusted association by sex and age between some FTO genes and changes of HDL-C
overtime are signi cant. Our ndings provide basis for searching for genes affecting change in HDL-C.
KEY WORDS: HIGH DENSITY LIPOPROTEIN CHOLESTEROL, GENETIC INFLUENCE, LATENT GROWTH CURVE
236
ARTICLE INFORMATION:
*Corresponding Author: moghimb@yahoo.com
Received 17
th
Dec, 2016
Accepted after revision 17
th
March, 2017
BBRC Print ISSN: 0974-6455
Online ISSN: 2321-4007 CODEN: USA BBRCBA
Thomson Reuters ISI ESC and Crossref Indexed Journal
NAAS Journal Score 2017: 4.31 Cosmos IF : 4.006
© A Society of Science and Nature Publication, 2017. All rights
reserved.
Online Contents Available at: http//www.bbrc.in/

Mahdi Akbarzadeh et al.
BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS SIMULATION STUDY OF CONTROLLING WATER CONING IN OIL RESERVOIRS 237
INTRODUCTION
High-density lipoproteins (HDL) are one of the ve
major groups oflipoproteins. Lipoproteins are complex
particles composed of multiple proteins which trans-
port all fat molecules (lipids) around the body within
the water outside cells. They are typically composed
of 80-100 proteins per particle and transporting up
to hundreds of fat molecules per particle. Unlike the
larger lipoprotein particles which deliver fat molecules
to cells, HDL particles remove fat molecules from cells
which need to export fat molecules. The fats carried
include cholesterol, phospholipids, and triglycerides;
amounts of each are quite variable. Increasing concen-
trations of HDL particles are strongly associated with
decreasing accumulation of atherosclerosis within the
walls of arteries. This is important because atherosclero-
sis eventually results insudden plaque ruptures,cardio-
vascular disease,strokeand othervascular diseases. HDL
particles are sometimes referred to as “good cholesterol”
because they can transport fat molecules out of artery
walls, reduce macrophage accumulation, and thus help
prevent or even regress atherosclerosis. However, studies
have shown that HDL-lacking mice still have the ability
to transport cholesterol to bile, suggesting that there are
alternative mechanisms for cholesterol removal. Also
heritability of low HDL-C is demonstrated, (Asselbergs
etal. 2012). Qureshi etal. have shown FTO genes have
association with HDL-C in Pakistani people, (Qureshi
etal. 2016).
In both epidemiological and clinical studies, as well
as the meta-analyses thereof, low plasma levels of high-
density lipoprotein (HDL) cholesterol (HDL-C) identi ed
individuals at increased risk of major coronary events.
In line with a causally protective effect, HDLs exert a
broad spectrum of potentially anti-atherogenic proper-
ties. Moreover, atherosclerosis was decreased or even
reverted in several animal models by transgenic over-
expression or exogenous application of apolipoprotein
(apoA-I) the most abundant protein of HDL,(Zhang etal.
2008).
However, to date, drugs increasing HDL-C, such as
brates, niacin, and inhibitors of cholesteryl ester trans-
fer protein (CETP), have failed to consistently and signif-
icantly reduce the risk of major cardiovascular events,
especially when combined with statins. Moreover, muta-
tions in several human genes as well as targeting of
several murine genes modulate HDL-C levels without
changing cardiovascular risk and atherosclerotic plaque
load, respectively, in the opposite direction as expected
from the inverse correlation of HDL-C levels and car-
diovascular risk in epidemiological studies. Because of
these controversial data, the pathogenic role and, hence,
suitability of HDL as a therapeutic target has been
increasingly questioned. Because of the frequent con-
founding of low HDL-C with hypertriglyceridemia, it has
been argued that low HDL-C is an innocent bystander
of (postprandial) hypertriglyceridemia or another culprit
related to insulin resistance or in ammation, (Gordon
etal. 1977; Silventoinen etal. 2007). Genetic associa-
tion of HDL-C is demonstrated in some population,(Kim
& Lee 2013; Nirengi etal. 2016; Bandarian etal. 2016).
In the Framingham study, high density lipoprotein
is shown as a protective factor against coronary heart
disease,(Gordon et al. 1977). In some studies on Ira-
nian population, low HDL-C is reported as a risk fac-
tor of coronary artery disease(Shari etal. 2009; Shari
et al. 2008; Hatmi et al. 2007).Also Akbarzadeh et al.
have shown some genes association with HDL-C in the
Iranian people without any results about the trend of
HDL-C(Akbarzadeh etal. 2011; Alavi Majd etal. 2010).
Among Iranian people as well as Tehranian people the
topic has been proven (Zarkesh etal. 2012).
An advanced, powerful and exible framework to
model the latent variables is structural equation mod-
eling (SEM). In recent years, in genetic analysis of longi-
tudinal family data, SEM has made signi cant progress.
One of these models is Morris and his colleagues that it
would be in addition to the relationships between vari-
ables over time, taking into account the simultaneous
equations to analyze genetic family data(Morris et al.
2010). The model can be able to run a wide range of
genetic models with latent variables under SEM, and in
recent months the model R package, strum, has been
released(Song etal. 2015).
According to our knowledge, up to this point about
the relationship between HDL-C changes related genes
in Iran has not been investigated. The aim of this study
was to assess the association of SNP in the gene fat mass
and HDL-C-associated gene (FTO) with HDL-C change
in a subset of participated families in Tehran Lipid and
Glucose Study (TLGS). To achieve this target we used the
latent growth curve model (LGCM) in STRUM R package
in TLGS family data.
MATERIAL AND METHODS
Population and sample: The TLGS is an ongoing longitu-
dinal large-scale community-based study, with a 3-year
follow-up period, designed to estimate the prevalence
of non-communicable disorders (NCD) and included a
representative sample of residents of 13 districts of Teh-
ran, capital of Iran. The TLGS has been implemented in
a multistage strati ed (district) cluster (families) random
sampling technique, to select more than 15000 people
aged > 3 years, from March 1999 to December 2011.The
phases II, III, and IV were prospective follow-up studies
Mahdi Akbarzadeh et al.
238 GENETIC ASSOCIATION ON HIGH-DENSITY LIPOPROTEINS CHANGES BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS
and were performed from 2002 to 2004, 2006 to 2008,
and 2009 to 2011, respectively, (Azizi etal. 2002; Azizi
et al. 2009). A total of 14761 individuals (valid case)
were selected from the total participating cases in TLGS,
including 3980 families, with an average number of 3.38
individuals, among the phase I (baseline).
This family-based study was conducted on families
who participated in TLGS, with at least two members
affected with syndrome metabolic (Mets).The design of
TLGS includes two major components, a cross-sectional
prevalence study of CVD and associated risk factors and
a prospective 20-year follow up in ve phases(14). In
the study 914 individual including 126 families with size
8.49±3.10 person from participated people in TLGS. Par-
ticipants provided informed consent and the study was
approved by the institutional ethics committees of the
Research Institute for Endocrine Sciences af liated to
Shahid Beheshti University of Medical Sciences, Tehran,
Iran. Information on age, sex, demographic and medica-
tion usage for treatment diabetes, hypertension and lipid
disorders were collected with a standardized question-
naire.
SNP Selection and Typing: Genomic DNA was
extracted from peripheral blood using standard salting-
out method(Koshy etal. 2016). 5 selected polymorphisms
(rs708272, rs1864163, rs1558902, rs7202116, rs9939609)
studied by Tetra ARMS method, by determining through
the NCBI site. MAF among for the ve markers is mini-
mum 0.039 and maximum 0.363. Which is as follows:
Our T-ARMS assay with different inner allele speci c
primers to produce allele-speci c PCR products. Two
outer primers will produce a PCR product to be used as
an internal control for reaction. For all studied SNPs,
the PCR reaction was optimized in a 12.5 μl total vol-
ume containing 1.5 μ DNA template, 6.25 μl Master Mix
containing (MgCl2, Smart Taq polymerase (Cinna Gene
Co; Iran) and BSA 0.1% (TaKaRa; Japan) and 2 μl primer
containing (outers and inners ) and 2.75 μl water.
Details of the primers information and nal frag-
ments are mentioned in table 1. The optimized thermo
cycling conditions were as follows: 5 min at 95 °C; 10
cycles of 45 sec at 94°C, 30 sec at 63°C, and 35 sec at
72°C; followed by 25 cycles of 45 sec at 94°C, 30 sec at
61°C, and 30 sec at72°C; and nal extension at72°C for 5
minutes. When the PCR products were separated by size
via agarose gel electrophoresis, each genotype generated
a special band. Accuracy of results was con rmed by
direct sequencing of 10% samples using outer primers.
Statistical Model: An advanced, powerful and ex-
ible framework to model the latent variables is structural
equation modeling (SEM).Structural equation modeling
evaluates the relationships among manifest variables,
manifest-latent variables and latent variables. Moreo-
ver the latent growth curve modeling (LGCM) has been
recently developed in longitudinal study (Bollen& Cur-
ran 2006). One of advantage of the model is that allows
for individual changes to be analyzed in the SEM frame-
work and includes additional latent factors in the lin-
ear model for testing a linear, quadratic, cubic, or spline
growth trajectory (Xitao & Xiaotao 2005).One advan-
tage of this model compared to the routine analytical
methods for longitudinal data is that modeling latent
variables over time and also the intercept and slop mod-
eled as latent variables.
The recent presented SEM is capable to model a broad rela-
tionships between latent variables based linkage or association
analysis. In strum a novel score test developed. This method
is a computationally rapid test of association with many SNPs
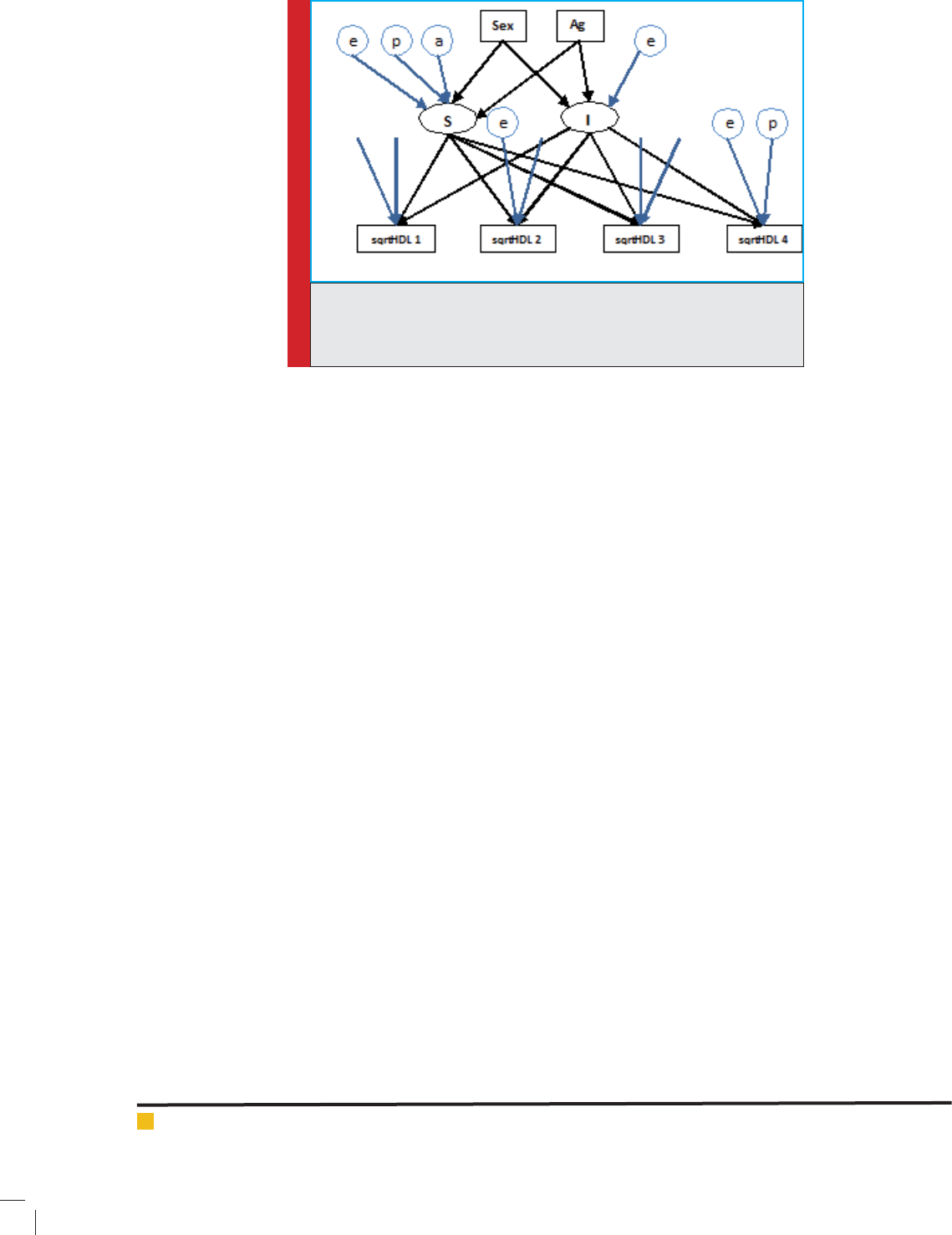
FIGURE 1. The path diagram of the latent growth curve model p: polygenic
effects; e: random effects; I: intercept; S: slope; sqrtHDL i: Squared meas-
urement of the HDL-C at ith time point; sex and age: gender and age of
individuals.
Mahdi Akbarzadeh et al.
BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS GENETIC ASSOCIATION ON HIGH-DENSITY LIPOPROTEINS CHANGES 239
in GWAS data (Song etal. 2015). In this new score test, we
rst assessed the null model t without any SNPs in the model
to con rm the appropriateness of the model for the data. In
the research, HDL-C measured in 4 time point. We examined
association of 5 markers of FTO genes with change of HDL-C
in Tehranian pedigrees using strum R package.
The graphical representations of analysis model with
latent variable for longitudinal HDL-C are shown for the
latent growth curve model in gure 1. Variable sqrtHDL-
C is the main trait values corrected for the use of nor-
mality. Nodes marked with: “p” are polygenic effects,
and “e” are random effects. “I” is the intercept and S is
the slope.
After removing the SNPs with no variation in the data
set or with no score test results, the remaining 18 SNPs
were tested for association with the main trait by coding
additively as 0, 1, or 2 based on the minor allele count.
RESULTS AND DISCUSSION
The genomic in ation factors were 1.01 and 1.02 for
each model. The MAF (minor allele frequency) of all
SNPs are greater than 5% and they are informative for
association analysis. Also the CFI (comparative t index)
of the conceptual model was 99% shown that the model
had good tness. The association between 5 FTO markers
and slop of HDL-C was signi cant (ie, P value<1.0e-5).
Table 1 shows the characteristics and P values.
Changing in HDL-C in human is certainly under
genetic in uence(Kim & Lee 2013; Nirengi etal. 2016;
Bandarian etal. 2016). Actually, the changes of HDL-C
is a latent variable. A preferable understanding way to
the change discovery is measuring by natural indica-
tor such as HDL-C. The study is the rst to simultane-
ously analyze level and change in HDL-C using a popu-
lation-based longitudinal study of individual including
families. We found a sigini cant genetic in uence on
change in HDL-C for four known markers. The modeling
assumes that long-term (15 year) individual HDL-C tra-
jectories in the four waves of young adults aged 3–83
years at baseline is adequately described by individual
initial level and rate of change.
CONCLUSION
Our ndings provide basis for searching for genes
affecting change in HDL-C. We assume that the genetic
in uence on change in HDL-C has a polygenic origin
with nite known markers. However, trying to identify-
ing genetic variants for HDL-C change may be require
a whole human genome wide scan in the target popula-
tion.
ACKNOWLEDGMENTS
We express appreciation to the participations and TLGS
staff for their guidance in the preparing dataset. We
thank the reviewers for their insightful and helpful com-
ments.
REFERENCES
Akbarzadeh, Mahdi, H A Majd, Maryam Sadat Daneshpour,
Yadolah Mehrabi, and Freydoon Azizi. (2011). Analyzing of
Multivariate Two Levels Haseman-Elston Regression and Its
Application in Genetic Linkage of HDL-C, Triglycerides and
Waist in 91 Iranian Families with Metabolic Syndrome. Koo-
mesh 12 (3). Semnan University of Medical Sciences and
Health Services: Pe266-Pe271.
Alavi Majd, H, M Akbarzadeh, Y Mehrabi, M S Daneshpour,
and F Azizi. (2010). Genetic Linkage Analysis of HDL-C in 91
Iranian Families with Metabolic Syndrome with Haseman-
Elston Regression Methods. HBI_Journals 7 (4): 243–47.
Asselbergs, Folkert W, Yiran Guo, Erik P A van Iperen, Suthesh
Sivapalaratnam, Vinicius Tragante, Matthew B Lanktree, Leslie
A Lange, Berta Almoguera, Yolande E Appelman, and John
Barnard. (2012). Large-Scale Gene-Centric Meta-Analysis
across 32 Studies Identi es Multiple Lipid Loci. The American
Journal of Human Genetics 91 (5). Elsevier: 823–38.
Azizi, Fereidoun, Arash Ghanbarian, Amir Abbas Momenan,
Farzad Hadaegh, Parvin Mirmiran, Mehdi Hedayati, Yadollah
Mehrabi, and Saleh Zahedi-Asl. (2009). Prevention of Non-
Communicable Disease in a Population in Nutrition Transition:
Tehran Lipid and Glucose Study Phase II. Trials 10 (January):
5. doi:10.1186/1745-6215-10-5.
Azizi, Fereidoun, Maziar Rahmani, Habib Emami, P Mirmiran,
R Hajipour, M Madjid, J Ghanbili, (2002). Cardiovascular Risk
Factors in an Iranian Urban Population: Tehran Lipid and Glu-
cose Study (Phase 1). Sozial- Und Präventivmedizin 47 (6).
Birkhäuser Verlag: 408–26. doi:10.1007/s000380200008.
Bandarian, Fatemeh, Maryam Sadat Daneshpour, Mehdi
Hedayati, Mohsen Naseri, and Fereidoun Azizi. (2016). Identi-
cation of Sequence Variation in the Apolipoprotein A2 Gene
and Their Relationship with Serum High-Density Lipoprotein
Cholesterol Levels.Iranian Biomedical Journal 20 (2). Pasteur
Institute of Iran: 84.
Bollen, KA, and PJ Curran.( 2006). Latent Curve Models: A
Structural Equation Perspective. John Wiley & Sons, Inc.
Tab le 1. SNPs associated with HDL-C in
analysis models
SNP Al MAF P value
rs708272 G/T 0.1232 3.320E-16
rs1558902 G/T 0.1102 2.045E-13
rs7202116 C/T 0.1851 1.489E-06
rs9939609 T/T 0.1448 2.276E-06
rs6696438 C/T 0.1476 4.462E-06
Al: major/minor alleles; MAF: minor allele frequency

Mahdi Akbarzadeh et al.
240 GENETIC ASSOCIATION ON HIGH-DENSITY LIPOPROTEINS CHANGES BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS
Gordon, Tavia, William P Castelli, Marthana C Hjortland, Wil-
liam B Kannel, and Thomas R Dawber. (1977). High Density
Lipoprotein as a Protective Factor against Coronary Heart Dis-
ease: The Framingham Study. The American Journal of Medi-
cine 62 (5). Elsevier: 707–14.
Hatmi, Z N, S Tahvildari, A Gafarzadeh Motlag, and A Sabouri
Kashani. (2007). Prevalence of Coronary Artery Disease Risk
Factors in Iran: A Population Based Survey BMC Cardiovascu-
lar Disorders 7 (1). BioMed Central: 1.
Kim, Jongwoo, and Seon Yeong Lee (2013 )Obesity and High-
Density Lipoprotein Cholesterol (HDL-C): The Recent Related
Research Trend Including New Generation Drugs for HDL-C
The Korean Journal of Obesity 22 (2): 67–72.
Koshy, Linda, A L Anju, S Harikrishnan, V R Kutty, V T Jissa,
Irin Kurikesu, Parvathy Jayachandran, A Jayakumaran Nair, A
Gangaprasad, and G M Nair (2016). Evaluating Genomic DNA
Extraction Methods from Human Whole Blood Using End-
point and Real-Time PCR Assays. Molecular Biology Reports.
Springer, 1–12.
Morris, Nathan J, Robert C Elston, and Catherine M Stein. (2010).
A Framework for Structural Equation Models in General Pedi-
grees. Human Heredity 70 (4): 278–86. doi:10.1159/000322885.
Nirengi, Shinsuke, Mami Fujibayashi, Kokoro Tsuzaki, Sachiko
Furuno, Akihiko Uchibe, Yasuharu Kawase, Kazuhiko Kotani,
and Naoki Sakane. (2016) ACTN3 Gene R577X Polymorphism
Associated with High-Density Lipoprotein Cholesterol and
Adiponectin in Rugby Players. Endocrine Practice. American
Association of Clinical Endocrinologists.
Qureshi, Sarah Anwar, Amir Mumtaz, and Saleem Ullah Sha-
hid. (2016)A Common Variant, rs3751812, in Fat Mass and
Obesity Associated (FTO) Gene Is Associated with Serum High
and Low Density Lipoprotein Cholesterol in the Pakistani Sub-
jects. Nutrition. Elsevier.
Shari , F1, S N Mousavinasab, M Saeini, and M Dinmoham-
madi. (2009). Prevalence of Metabolic Syndrome in an Adult
Urban Population of the West of Iran. Experimental Diabetes
Research 2009. Hindawi Publishing Corporation.
Shari , F, S N Mousavinasab, R Soruri, M Saeini, and M Din-
mohammadi (2008) High Prevalence of Low High-Density
Lipoprotein Cholesterol Concentrations and Other Dyslipi-
demic Phenotypes in an Iranian Population Metabolic Syn-
drome and Related Disorders 6 (3). Mary Ann Liebert, Inc. 140
Huguenot Street, 3rd Floor New Rochelle, NY 10801-5215 USA:
187–95.
Silventoinen, Karri, Meike Bartels, Daniëlle Posthuma, G Fred-
eriek Estourgie-van Burk, Gonneke Willemsen, Toos C E M van
Beijsterveldt, and Dorret I Boomsma (2007) Genetic Regulation
of Growth in Height and Weight from 3 to 12 Years of Age:
A Longitudinal Study of Dutch Twin Children Twin Research
and Human Genetics 10 (2). Cambridge Univ Press: 354–63.
doi:https://doi.org/10.1375/twin.10.2.354.
Song, Yeunjoo E, Catherine M Stein, and Nathan J Mor-
ris. (2015) Strum: An R Package for Structural Modeling of
Latent Variables for General Pedigrees.BMC Genetics 16 (1):
35. doi:10.1186/s12863-015-0190-3.
Xitao, Fan, and Fan Xiaotao (2005) Power of Latent Growth
Modeling for Detecting Linear Growth: Number of Measure-
ments and Comparison with Other Analytic Approaches. The
Journal of Experimental Education 73 (2). Taylor & Francis:
121–39.
Zarkesh, Maryam, Maryam Sadat Daneshpour, Bita Faam,
Mohammad. (2012) Heritability of the Metabolic Syndrome
and Its Components in the Tehran Lipid and Glucose Study
(TLGS). Genetics Research 94 (6): 331–37. doi:10.1017/
S001667231200050X.
Zhang, Da-Wei, Rita Garuti, Wan-Jin Tang, Jonathan C Cohen,
and Helen H Hobbs (2008) Structural Requirements for PCSK9-
Mediated Degradation of the Low-Density Lipoprotein Recep-
tor Proceedings of the National Academy of Sciences 105 (35).
National Acad Sciences: 13045–50.