Biomedical
Communication
Biosci. Biotech. Res. Comm. 10(1): 34-43 (2017)
Machine learning approaches to study HIV/AIDS
infection: A Review
Sweta Kumari, Usha Chouhan and Sunil Kumar Suryawanshi*
Department of Bioinformatics, Maulana Azad National Institute of Technology (MANIT), Bhopal- 462051
(M.P.), INDIA
ABSTRACT
In this review, PubMed database has been explored to elucidate the problems related to HIV/AIDS, which have
been solved previously using various machine learning approaches and some other techniques. Literatures from the
epidemic years of HIV/AIDS till February, 2017 have been examined and problems such as prediction of HIV/AIDS
protease cleavage sites and inhibitors, prediction of coreceptors usage for viral entry, development of anti-viral
agents and prediction of response, resistance and adverse effect of antiretroviral therapy have been considered for
the current study. Complications associated with HIV/AIDS infection as well as all three stages of HIV infection have
been described. HIV virus binding to the coreceptors CCR5 and CXCR4 are delineated to show the signi cant role of
the coreceptors for the anti-HIV drug development. After exploring various datasets, viral tropisms are found to be
relevant to the viral third V3 region of the HIV virus binding.
KEY WORDS: HIV, MACHINE LEARNING TECHNIQUES, V3 REGION, CD4 RECEPTOR, CCR5 AND CXCR4 CORECEPTORS
34
ARTICLE INFORMATION:
*Corresponding Author: bioinfosunil@gmail.com
Received 29
th
Dec, 2016
Accepted after revision 3
rd
March, 2017
BBRC Print ISSN: 0974-6455
Online ISSN: 2321-4007 CODEN: USA BBRCBA
Thomson Reuters ISI ESC and Crossref Indexed Journal
NAAS Journal Score 2017: 4.31 Cosmos IF : 4.006
© A Society of Science and Nature Publication, 2017. All rights
reserved.
Online Contents Available at: http//www.bbrc.in/
INTRODUCTION
Human immunode ciency virus/acquired immune de -
ciency syndrome (HIV/AIDS) was originated from mon-
key in the United States in 1981. AIDS is a chronic and
potentially most threatening infectious disease caused
by human immunode ciency virus in the 21
st
century.
78 million people were estimated to be suffering from
HIV/AIDS and 35 million people have died since the
start of the epidemic year but 36.7 million people were
reported as HIV/AIDS infected and 1.1 million people
have died in 2015 globally; 2.1 million people were
found to be newly HIV/AIDS infected globally (Alkema
2016). Eastern and Southern Africa have a maximum
increase since almost the start of the epidemic year. HIV/
AIDS regularly decimated the population of the Africa
shown in gure1 (HIV/AIDS 2016).
The current prevalence of the HIV/AIDS is 0.8% of
the worldwide. 18.2 million people were accessing
antiretroviral therapy in June 2016 (Alkema et al. 2016).

Sweta, Usha and Sunil
Antiretroviral therapy (ART) has a very crucial role in
the HIV treatment, but HIV infected individuals acquire
resistance to the ART over a certain period. Antiretrovi-
ral therapy coverage increased from 7.51 million in 2010
to 17.03 million in 2015 shown in gure 2.
However, scientists are working to develop one vac-
cine to prevent HIV/AIDS. NIH supported clinical trial
that was launched in 2016 to test a possible HIV/AIDS
vaccine. This vaccine trial HVTN 072 is testing whether an
experimental vaccine regimen safely prevents HIV/AIDS
infection among South African adults (Health 2015).
The determination of the 90-90-90 treatment target
is to be achievable by the reinforcement of the con-
tinuing momentum in 2020, whereby 90% of the HIV/
AIDS infected people aware of their HIV/AIDS status,
90 % of the people with known their HIV/AIDS positive
status are taking treatment and 90% of the people on
HIV/AIDS treatment have suppressed viral loads (NAS-
COP 2014). Recent updated UNAIDS estimates indicate
that US$ 26.2 billion will be required for the HIV/AIDS
response in 2020, with US$ 23.9 billion required in 2030
(NASCOP 2014). The world has committed to ending the
HIV/AIDS epidemic by 2030 (Bernard 2016).
METHODS
We searched PubMed database with the keywords of
“HIV” and “Machine Learning Approaches” up to Feb-
ruary 22, 2017 and starting time was not given. The
number of the articles retrieved from the PubMed data-
base was 114, out of which there was 6 review articles.
Efforts here made to review some uncertainties related
to HIV infection. The search completeness was examined
by the list of references of the reviewed articles. Some-
where, Book chapters were also included, some articles
were excluded on criteria such as articles not in English
language, campaign posters, newspaper articles and the
articles irrelevant to the topic of this review. In numbers,
32 originally related articles were selected on the basis
of this evaluation are listed in Table-1.
FIGURE 1. New HIV infections among people aged 15 years and above in different
regions since last six years from 2010 to 2015(Alkema et al. 2016)
FIGURE 2. Antiretroviral therapy coverage % in different regions since last six
years from 2010 to 2015(Bernard 2016)
BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS MACHINE LEARNING APPROACHES TO STUDY HIV/AIDS INFECTION 35

Sweta, Usha and Sunil
STAGES OF HIV/AID
There are mainly three stages of HIV: Primary infection
(Acute HIV), Clinical latent infection (Chronic HIV) and
Early symptomatic HIV infection (Wawer 2005). In pri-
mary infection, a u-like illness developed within one or
two months after the virus entering the body of people,
including signs and symptoms such as fever, headache,
muscle aches, joint pain, rash, sore throat and swollen
lymph glands mainly on the neck. Though, the symp-
toms of the rst stage of HIV infection are mostly unno-
ticed, the amount of virus in the viral load or bloodstream
spreads very highly at this time, resulting in dispersing
the HIV infection more ef ciently during primary infec-
tion than the next stage. In the clinical latent infection,
generally persistent swelling of lymph nodes occurs and
HIV remains in the human body but with no signs and
symptoms. This stage lasts around 10 years in people not
receiving antiretroviral medications and lasts for decades
in people receiving antiretroviral therapy. Some people
expedite to more intense disease much sooner. In the last
stage of early symptomatic HIV infection, people probably
progress to mild infections or chronic signs and symp-
toms such as fever, fatigue, swollen lymph nodes, diarrhea,
weight loss, oral yeast infection (thrush) and shingles (her-
pes zoster) due to continuous replication of the virus in the
human body and distortion of the immune cells.
COMPLICATIONS OF HIV/AIDS
The burden of HIV is to a large extent consequence of
infections (tuberculosis (TB), cytomegalovirus, candidi-
asis, cryptococcal meningitis, toxoplasmosis and crypto-
sporidiosis), cancers (Kaposi’s sarcoma and lymphomas)
and others (wasting syndrome, neurological complica-
tions and kidney dis How ease) (How). The increasing in
the incidences of the HIV/AIDS and their consequences
in terms of falling down the number of CD4 (Cluster of
differentiation 4) receptors and coreceptors lead to dam-
age the immune system of the human. Therefore, iden-
ti cation of the used coreceptor for the viral entry that
can target to block the coreceptor to bind with the virus
and maintenance of the CD4 receptors and coreceptors
is being crucial for new therapeutic agents for the treat-
ment of immune de ciency syndrome (Kaplan 2009).
Infections common to HIV/AIDS
Tuberculosis (TB): Tuberculosis coinfection is associated
with increase viral replications. TB is the most occurring
infection related to HIV/AIDS and a leading cause of death
(Pawlowski 2012).
Cytomegalovirus: This herpes virus is transmitted in
human body uids such as saliva, urine, blood, breast milk
and semen. The virus remains inactive in a healthy immune
system and remains resurfaces in weakens immune sys-
tem causing damage to eyes, lungs, digestive tract or other
organs (Mathevula 2013).
Candidiasis: Candidiasis causes in ammation and a thick,
white coating on the mucous membranes of mouth, tongue,
vagina or esophagus (Cutlan 2010).
Cryptococcal Meningitis: Cryptococcal meningitis is a
common infection of the central nervous system associated
with HIV/AIDS, caused by a fungus found in soil. Menin-
gitis is an in ammation of the membranes and uid sur-
rounding brain and spinal cord (Park 2009).
Toxoplasmosis: This infection is caused by Toxoplasma
gondii, a parasite dispersed primarily by cats. Infected
cats successfully pass the parasites in their stools, and the
parasites may then transfer to other animals and humans
(Berger-Schoch 2011).
Cryptosporidiosis: An intestinal parasite usually found
in animals and it is responsible for the infection. Crypto-
sporidiosis contracted when human ingests contaminated
food or water. The parasite grows in the intestines of human
and bile ducts, leading to severe, chronic diarrhea in HIV/
AIDS infected people (Mathewos 2014).
Cancers common to HIV/AIDS
Kaposi’s sarcoma: Kaposi’s sarcoma is a blood vessel walls,
a very rare tumor in HIV-negative people but a very com-
mon in HIV-positive people. It usually appears as pink, pur-
ple or red lesions on the skin and mouth of the HIV infected
people (Page 2006). Internal organs can also be affected by
Kaposi’s sarcoma (Di Benedetto 2008).
Lymphomas: Lymphomas cancer originates in white blood
cells and usually appears in lymph nodes. Painless swelling
of the lymph nodes in neck, armpit or groin is most com-
mon early sign (Daniels 2012).
Other complications
Wasting syndrome: Aggressive treatment regimens have
decreased the number of cases of wasting syndrome, but
it still affects many HIV/AIDS infected people. It leads to
a loss of at least 10 percent of human body weight, often
escorted by diarrhea, chronic weakness and fever (Bass
2015).
Neurological complications: It can cause neurological
symptoms such as confusion, depression, forgetfulness,
anxiety and dif culty walking, whereas AIDS doesn’t
infect the nerve cells. AIDS dementia complex is one of
the most common neurological complications, which leads
to changes in behavior and diminishes in mental function
(Ances 2007).
Kidney disease: HIV-associated nephropathy (HIVAN) is an
in ammation of the small lters in kidneys that remove
extra uid and wastes from bloodstream and pass them to
your urine. The risk of developing HIVAN is much higher
in black people due to a genetic predisposition. A antiret-
roviral therapy should be started in those diagnosed with
HIVAN in any case of CD4 count (Scherzer 2012).
36 MACHINE LEARNING APPROACHES TO STUDY HIV/AIDS INFECTION BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS

Sweta, Usha and Sunil
Receptors
CD4 receptors are particularly attractive target mol-
ecules, since they have been found to play a certain role
in maintaining the immune system of the human. CCR5
and CXCR4 are two coreceptors of the receptor present
on the T-cell, which is useful for viral cellular entry
shown in gure 3.
CCR5 (C-C chemokine receptor type 5)
The CCR5 protein belongs to the family of the beta
chemokine receptors of the integral membrane proteins
(Samson 1996). The CCR5 coreceptors are G protein-
coupled coreceptors (Aavikko 2014). CCR5 coreceptors
are also known as CD195 proteins expressed on the sur-
face of white blood cells, T helper cells, macrophages,
dendritic cells, eosinophils and microglia. T helper cells
are speci c tissues and organ targets by which HIV virus
causes AIDS using CCR5 coreceptors to enter and infect
into the immunological cells. The viral entry of HIV-1
into a target host cell is enabled by the essential HIV-1
envelope glycoprotein structure (Alkhatib 2009). The
Gp120 external subunit and Gp41 transmembrane subu-
nit are two subunits of the envelope glycoprotein struc-
ture cleaved from a Gp160 protein precursor encoded by
the HIV-1 env gene (Checkley 2011).
The Gp120 subunit binds to a CD4 glycoprotein and
a HIV-1 coreceptor CCR5 expressed on a target host cell
forming a heterotrimeric complex (Murphy 2001). The
binding of Gp120 envelope protein to CCR5 coreceptor
consists two crucial steps: The tyrosine sulfated amino
terminus of this coreceptor is an “essential determinant”
of binding to Gp120 in the rst step and there must be
reciprocal action (synergy, intercommunication) between
gp120 and the CCR5 transmembrane domains in a sec-
ond step (Ji 2007). Some individuals lead to a muta-
tion as CCR5-delta 32 in the CCR5 gene by the genetic
deletion of a portion of the CCR5 gene to protect them
against these HIV strains. This mutation in homozygous
carriers is resistant to the M-tropic strains of the HIV-1
infected individuals (De Silva 2004 Hütter 2009 Allers
2011 Zhen 2013 Kay 2014 and Tebas 2014).
The CCR5 gene encodes the CCR5 protein, which is
located on the short (p) arm at the position of 21 on chro-
mosome 3.The cognate ligands of CCR5 include CCL3,
CCL4 and CCL3L1 (Struyf 2001 and Miyakawa 2002) and
CCR5 interacts with CCL5 (Struyf et al. 2001 Miyakawa
et al. 2002 and Slimani 2003).The general ligands for the
receptor RANTES, MIP-1 and MIP-1 are able to sup-
press HIV-1 infection in vitro (Nakayama 2012).
CXCR4 (C-X-C chemokine receptor type 4)
CXCR4 is an alpha-chemokine receptor speci c for
stromal-derived-factor-1 (SDF-1) also known as CD184
(cluster of differentiation 184) proteins. SDF-1 is a mol-
ecule endowed with effective chemotactic activity of
lymphocytes. During the implantation and hormone
replacement therapy cycle in the endomentrium, pro-
ducing, in the presence of human blastocyst and a sur-
face polarization of the CXCR4 coreceptors suggest that
these coreceptors are connected in the adhesion phase
of the human implantation, CXCR4 is upregulated. The
ligand of CXCR4, SDF-1 is known to be signi cant in
hematopoietic stem cell homing to the bone marrow and
in hematopoietic stem cell quiescence. CXCR4 signaling
coreceptors found to regulate the CD20 expression on B
cells (Pavlasova 2016).
SDF-1 and CXCR4 were believed to be a relatively
monogamous ligand-receptor pair (other chemokines are
promiscuous, tending to use several different chemokine
receptors) until recently. Recent evidence demonstrates
ubiquitin is also a natural ligand of CXCR4 (Saini 2010).
Ubiquitin is a small (76-amino acid) protein highly con-
served among eukaryotic cells. It is best known for its
intracellular role in targeting ubiquitylated proteins for
degradation via the ubiquitin proteasome system. Evi-
dence in numerous animal models suggests ubiquitin is
anti-in ammatory immune modulator and proin am-
matory damage endogenous opponent associated molec-
ular pattern molecules (Majetschak 2011). It is speculated
this interaction may be through CXCR4 mediated sign-
aling pathways. MIF is an additional ligand of CXCR4
(Bernhagen 2007).
CXCR4 plays a role in neurological guidance by pre-
senting in newly generated neurons during embryo-
genesis and adult life. The receptor levels decrease as
neurons mature. CXCR4 mutant mice have aberrant
neuronal distribution. This has been implicated in disor-
ders such as epilepsy (Bagri 2002).
The viral regions
Coreceptor consideration holds the V1-V2 region
oh Gp120 and the bridging sheet such as antiparallel
and 4-stranded sheet that joints the inner and outer
domains of the Gp120. The coreceptor usage through its
peptide composition and the degree of N-linked glyco-
sylation can be in uenced by the V1-V2 stem. In con-
trast with V1-V2 region, the V3 loop region is highly
mutable and therefore it is the most signi cant determi-
nant of coreceptor speci city (Bozek 2013).
The HIV enzymes role
The viral genome reverse transcription is required to
generate the proviral DNA and the integration into the
target genome cell for successful HIV-1 virus replication.
The viral encoded enzymes such as reverse transcriptase
(RT) and integrase (IN) put their potential for these
events and act sequentially during viral replication. The
BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS MACHINE LEARNING APPROACHES TO STUDY HIV/AIDS INFECTION 37

Sweta, Usha and Sunil
FIGURE 3. HIV virus cellular entry through binding with the CD4 receptor and
CXCR4 and CCR5 coreceptor in T-tropic and M-tropic HIV respectively(Gorry
2005)
double stranded proviral DNA produced by replicating
both RNA and DNA templates using RT enzyme which
is a heterodimer of p66 and p51 subunits (Arts 1997).
A de ned set of DNA cleavage and combined events to
insert the proviral DNA into the host genome directed by
the IN enzyme which is 32 kDa polypeptide (Wiskerchen
1995). HIV-1 protease (PR) is important for the life-
cycle of the HIV virus and this enzyme is responsible
for cleavage of the newly synthesized polyproteins
at the appropriate places to create the mature protein
components of an infectious HIV virion. HIV virions
remain uninfected without effective HIV PR (Doitsh
2010).
The review highlights the scarcity of information on
the HIV/AIDS problems which are possible to solve using
various machine learning techniques are shortlisted in
Table 1. Classi cation of X4, R5 and dual (R5X4) tropic
HIV-1 from V3 loop amino acid sequences of HIV-1 sub-
type B using arti cial neural networks up to accuracy of
81.8% (Fogel et al. 2015) and classi cation of X4 and
R5tropic HIV-1 from V3 loop amino acid sequences of
HIV-1 subtype C using SVM showed a good concord-
ance of 85% (Gupta et al. 2015) are done in USA and
India respectively. Prediction of coreceptor usage for
viral entry from Gp120 V3 loop amino acid sequences
using SVM, heuristic and statistical learning methods,
two-level machine, random forest, boosted decision
tree, and neural network machine learning algorithms
has been done in India, Germany and USA respectively
(Raghava 2013 Sander et al. 2007 Dybowski 2010 and
Vaisman 2010). Prediction of HIV-1 protease cleavage
site and inhibitors using Feature selection subset method
of multi-layered perceptron (FS-MLP) learning, SVM,
ANN pharmacophore and docking methods have been
done 80.0% ~ 97.4% accuracy in Taiwan, Korea, Turkey
and China respectively (Kim et al. 2010 Singh Su 2016
Öztürk et al. 2013 and Wei et al. 2015).
Some of the literatures are regarding of drug devel-
opment, anti-viral agents development (Kirchmair et
al. 2011), antiretroviral response prediction (Zazzi et al.
2012 Prosperi 2011 and Prosperi et al. 2009), antiretro-
viral resistance prediction (Zazzi 2016 Riemenschneider
2016a Riemenschneider 2016b Heider et al. 2013 and
Kijsirikul 2008), antiretroviral adverse effects predic-
tion (Adrover et al. 2015) has been analyzed which were
used machine learning approaches, SVM, Expert’s Rules
and Linear and Non-linear statistical learning algo-
rithms, Radial basis function networks (RBF networks),
k-nearest neighbor (kNN) and Virtual screening method
to improve their result in Austria, Italy, USA, Germany
and Thailand. Prediction of antibody of HIV epitope net-
works using neutralization titers and a novel computa-
tional methods or a simple machine learning methods
has been done in USA (Evans et al. 2014 Hepler et al.
2014 and Choi et al. 2015.
Prediction of HIV-1 RT associated RNase H inhibition
(Poongavanam 2013) shown good enrichment (80-90%)
by receptor-based exible docking experiments com-
pared to ligand-based approaches such as FLAP (74%),
shape similarity (75%) and random forest (72%) in Den-
mark. Ligand based computational modeling studies on
non-nucleoside reverse transcriptase inhibitors of HIV-1
(Pancholi et al. 2014) has been done India respectively.
Prediction of bioactivities of HIV-1 integrase ST inhibi-
tors (Xuan et al. 2013) and classi cation of active and
weakly active ST inhibitors of HIV-1 integrase has (Yan
et al. 2012) been done using machine learning approaches
in China and USA respectively. Prediction of interactions
between HIV-1 and human proteins using SVM in China
has also done (Wei et al. 2015). One of the literatures has
worked on the detection of M tuberculosis in patients with
and without HIV coinfection by identifying a 251 gene
expression signatures using SVM with 81.4% and 88.9-
94.7% accuracy respectively in USA (Dawany et al. 2014).
38 MACHINE LEARNING APPROACHES TO STUDY HIV/AIDS INFECTION BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS
Sweta, Usha and Sunil
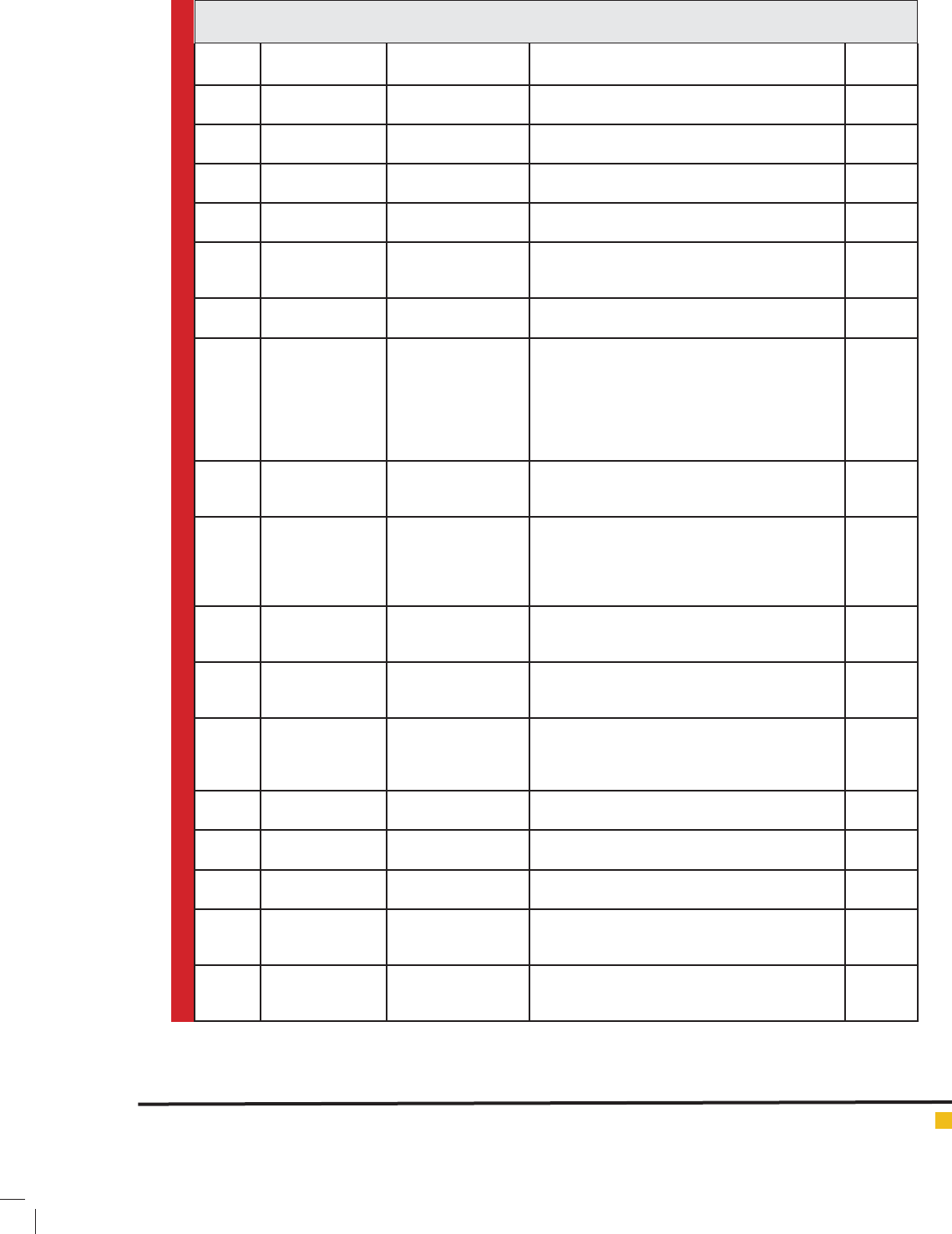
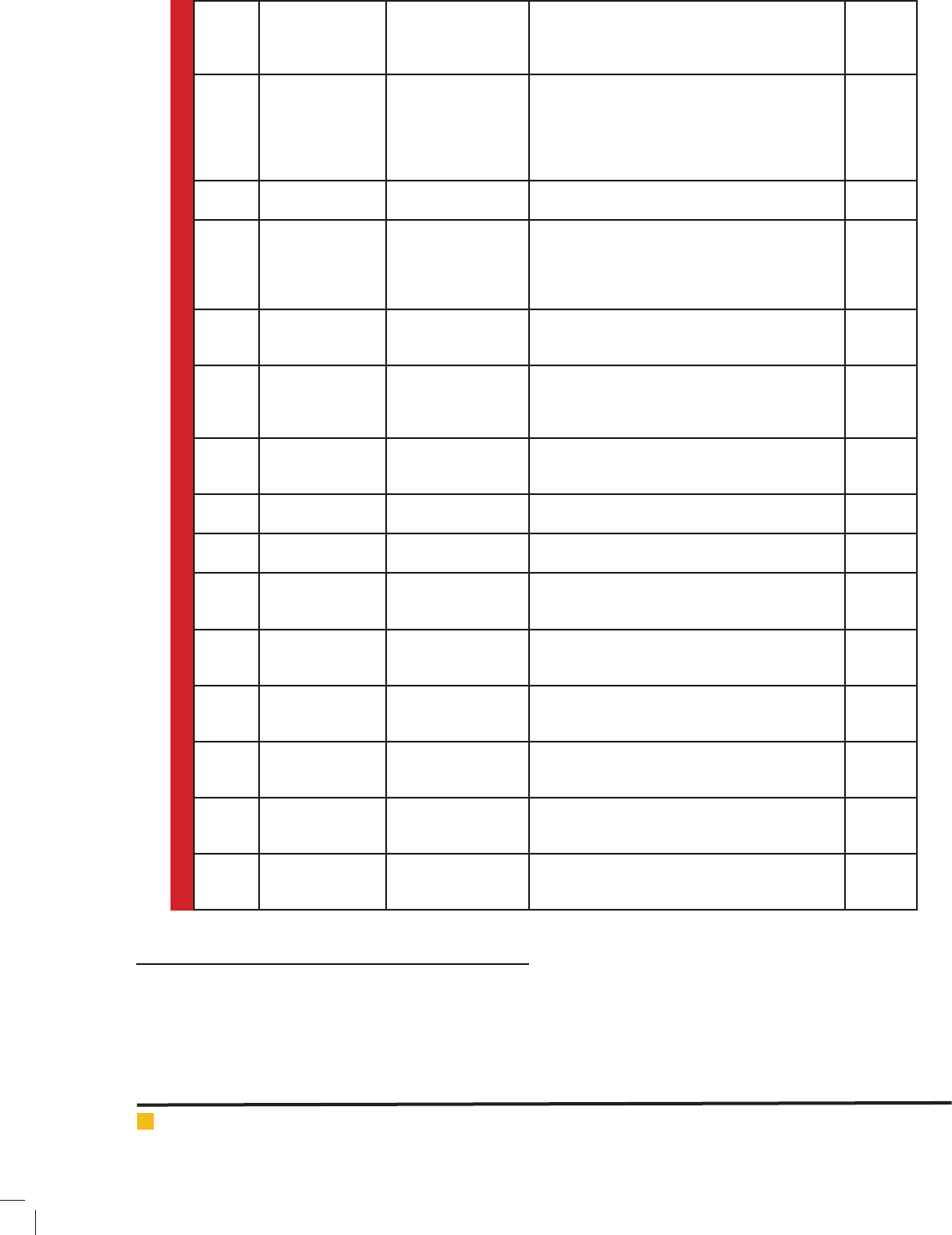
Table 1. Literatures involved for the current study for machine learning approaches used to solve various HIV/
AIDS problems.
Number Author name-
Publication year
Methods Objectives Country
1 (Gupta 2015) Machine learning
methods
Prediction of tropism in HIV-1 subtype C V3 loop
sequences using genotypic tools.
India
2 (Fogel 2015) Arti cial neural
networks
Classi cation of R5-, X4- and R5X4-tropics HIV-1
using evolved neural networks.
US
3 (Antell 2016) Machine learning Identi cation of R5- and X4-speci c Tat and LTR
sequence signatures using HIV envelope V3.
USA
4 (Kumar 2013) Support vector
machine
Prediction of HIV-1 coreceptor usages using hybrid
approach from its V3 loop amino acid sequences.
India
5 (Sander 2007) Heuristic and
Statistical learning
methods
Prediction of HIV-1 coreceptor usage by structural
descriptors of gp120 V3 loop.
Germany
6 (Dybowski 2010) Two-level machine
learning methods
Prediction of co-receptor usage of HIV-1 from
genotype.
Germany
7 (Masso 2010) Random forest,
support vector
machine, boosted
decision tree, and
neural network
machine learning
algorithms
Determination of HIV-1 co-receptor usage from
accurate and ef cient gp120 V3 loop structure based
models.
USA
8 (Evans 2013) Support vector
machine, PSSM and
11/25 rule
Genotypic prediction of HIV-1 coreceptor tropism
using a case-based reasoning system.
USA
9 (Kim 2010) Feature selection
subset method
of multi-layered
perceptron (FS-MLP)
learning
Analysis of HIV-1 protease cleavage site an MLP-
based feature subset selection.
Korea
10 (Singh 2016) Arti cial neural
networks
HIV-1 protease cleavage site prediction using
a combination of sequence, structural, and
physiochemical properties.
Taiwan
11 (Öztürk 2013) Support vector
machine
Prediction of HIV-1 protease cleavage site using a
consistency-based feature selection method allied
with linear SVMs.
Turkey
12 (Wei 2015) Support vector
machine, shape,
pharmacophore and
docking methods
Multistage virtual screening and novel identi cation
of HIV-1 protease inhibitors by integrating SVM,
shape, pharmacophore and docking methods.
China
13 (Kirchmair 2011) Virtual screening
techniques.
Development of anti-viral agents using molecular
modeling and virtual screening techniques.
Austria
14 (Zazzi 2012) Machine learning
approaches
Prediction of antiretroviral treatment response using
machine learning: The EuResist project.
Italy
15 (Prosperi 2011) Machine learning
approaches
Prediction of antiretroviral treatment response using
computational models.
USA
16 (Prosperi 2009) Linear and Non-linear
statistical learning
models
Prediction of response to antiretroviral treatment by
investigating of expert rule bases, logistic regression,
and non-linear machine learning techniques.
Italy
17 (Zazzi 2016) Expert’s Rules and
Statistical/Machine
learning algorithms.
Computer-Aided Optimization of Combined Anti-
Retroviral Treatment for HIV: New Drugs, Drug
Targets and Drug Resistance.
US
BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS MACHINE LEARNING APPROACHES TO STUDY HIV/AIDS INFECTION 39
Sweta, Usha and Sunil
18 (Riemenschneider
2016a)
Support vector
machine, Random
forest and Statistical
methods
Prediction of HIV Drug Resistance using Current
Computational Approaches.
Germany
19 (Riemenschneider
2016b)
Machine learning
techniques, binary
relevance classi ers,
classi er chains, and
ensembles of classi er
chains
Multiclass classi cation for HIV-1 drug resistance
prediction by exploiting cross-resistance information
with 662 protease sequences and 715 reverse
transcriptase sequences.
Germany
20 (Heider 2013) Machine learning
techniques.
Multiclass classi cation for HIV-1 drug resistance
prediction by exploiting cross-resistance information.
Germany
21 (Srisawat 2008) Support vector
machine, Radial basis
function Networks and
k-nearest neighbor
methods
Prediction of HIV-1 drug resistance by combining
classi ers.
Thailand
22 (Adrover et al.
2015)
Machine learning and
crowdsourced human
assessment
Identi cation of adverse effects of HIV drug
treatment and related sentiments using Twitter.
US
23 (Evans 2014) Boosted algorithms of
consisting of multiple
machine learnings and
statistical models
Prediction of HIV-1 broadly neutralizing antibody
epitope networks using neutralization titers and a
novel computational method.
USA
24 (Hepler 2014) Machine learning
methods
IDEPI: fast prediction of HIV-1 antibody epitopes
and other phenotypic features from sequence dataset
using a exible machine learning techniques.
USA
25 (Choi 2015) Machine learning
methods
Prediction of antibody feature: function relationships
in RV144 vaccines using machine learning methods.
USA
26 (Poongavanam
2013)
Virtual screening Prediction of HIV-1 RT related to RNase H inhibition
using virtual screening.
Denmark
27 (Pancholi 2014) SVM, Back
propagation neural
networks
Ligand based computational modeling studies on
non-nucleoside reverse transcriptase inhibitors of
HIV-1
India
28 (Xuan 2013) Support vector
machine and
Regression methods
Bioactivity of HIV-1 integrase ST inhibitors predicted
using multilinear regression analysis and support
vector machine.
China
29 (Yan 2012) Machine learning
methods
Support vector machine used for classi cation of
active and weakly active ST inhibitors of HIV-1
integrase.
USA
30 (Mei 2013) Support vector
machine
Prediction of interactions between HIV-1 and human
proteins using probability weighted ensemble transfer
learning.
China
31 (Dawany 2014) Support vector
machine
Detection of accurate M. tuberculosis in patients with
and without HIV co-infection by identi cation of a
251 gene expression signature.
USA
32 (Holman 2012) Machine learning
methods
Identi cation of amino acid signatures in the HIV
env gene predictive of dementia using a machine
learning approach.
USA
CONCLUSION
After reviewing all the literature, the coreceptor which
has been used for cellular viral entry is very necessary
to identify to develop the drugs that can target the core-
ceptor and prevent the coreceptor from binding with
HIV virus. Maraviroc has been found to be an as suc-
cessful barrier in blocking the CCR5 coreceptor but fails
in blocking the CXCR4 coreceptor. Prediction of HIV-1
coreceptor usage is necessary to identify the number of
CD4 counts remained into the body of HIV-1 infected
people. HIV-1 protease is a retroviral aspartyl protease
(retropepsin) that is essential for the HIV-1 lifecycle,
the retrovirus that is responsible for AIDS. The integral
40 MACHINE LEARNING APPROACHES TO STUDY HIV/AIDS INFECTION BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS

Sweta, Usha and Sunil
role of the HIV-1 protease in the viral replication, HIV-1
protease has become a prime target for drug therapy.
Identi cation of the coreceptor which has been used
for cellular viral entry with different machine learning
approaches and more high accuracy than the previous
ndings is our future work for contribution in the drug
development of HIV-1.
ACKNOWLEDGEMENT
We are thankful to the Department of Biotechnology
(DBT), New Delhi for providing support for this work
under Bioinformatics Infrastructure Facility of DBT at
Maulana Azad National Institute Technology, Bhopal.
REFERENCES
Aavikko, M. (2014) Identi cation of novel tumor predisposi-
tion families and underlying genetic defects.
Adrover, C., Bodnar, T., Huang, Z., Telenti, A. ,Salathé, M.
(2015) Identifying adverse effects of HIV drug treatment and
associated sentiments using Twitter. JMIR public health and
surveillance, Vol. 1, No. 2.
Alkema, L., Chou, D., Hogan, D., Zhang, S., Moller, A.-B.,
Gemmill, A., Fat, D.M., Boerma, T., Temmerman, M. ,Mathers,
C. (2016) Global, regional, and national levels and trends in
maternal mortality between 1990 and 2015, with scenario-
based projections to 2030: a systematic analysis by the UN
Maternal Mortality Estimation Inter-Agency Group. The Lan-
cet, Vol. 387, No. 10017, pp. 462-74.
Alkhatib, G. (2009) The biology of CCR5 and CXCR4. Current
Opinion in HIV and AIDS, Vol. 4, No. 2, p. 96.
Allers, K., Hütter, G., Hofmann, J., Loddenkemper, C., Rieger,
K., Thiel, E. ,Schneider, T. (2011) Evidence for the cure of HIV
infection by CCR5Δ32/Δ32 stem cell transplantation. Blood,
Vol. 117, No. 10, pp. 2791-9.
Ances, B.M. ,Ellis, R.J. (2007) Dementia and neurocognitive
disorders due to HIV-1 infection, Seminars in neurology, Vol.
27, Copyright© 2007 by Thieme Medical Publishers, Inc., 333
Seventh Avenue, New York, NY 10001, USA., pp. 086-92.
Antell, G.C., Dampier, W., Aiamkitsumrit, B., Nonnemacher,
M.R., Jacobson, J.M., Pirrone, V., Zhong, W., Kercher, K., Pas-
sic, S. ,Williams, J.W. (2016) Utilization of HIV-1 envelope V3
to identify X4-and R5-speci c Tat and LTR sequence signa-
tures. Retrovirology, Vol. 13, No. 1, p. 32.
Arts, E.J., Le Grice, S.F. (1997) Interaction of retroviral reverse
transcriptase with template–primer duplexes during replica-
tion. Progress in nucleic acid research and molecular biology,
Vol. 58, pp. 339-93.
Bagri, A., Gurney, T., He, X., Zou, Y.-R., Littman, D.R., Tessier-
Lavigne, M. ,Pleasure, S.J. (2002) The chemokine SDF1 reg-
ulates migration of dentate granule cells. Development, Vol.
129, No. 18, pp. 4249-60.
Bass, L.H., Washington, C.M. (2015) Infection Control in Radia-
tion Oncology Facilities. Principles and Practice of Radiation
Therapy, p. 178.
Berger-Schoch, A., Herrmann, D., Schares, G., Müller, N., Ber-
net, D., Gottstein, B. ,Frey, C. (2011) Prevalence and genotypes
of Toxoplasma gondii in feline faeces (oocysts) and meat from
sheep, cattle and pigs in Switzerland. Veterinary parasitology,
Vol. 177, No. 3, pp. 290-7.
Bernard, E., Cameron, S. (2016) Advancing HIV justice 2.
Building momentum in global advocacy against HIV crimi-
nalisation.
Bernhagen, J., Krohn, R., Lue, H., Gregory, J.L., Zernecke, A.,
Koenen, R.R., Dewor, M., Georgiev, I., Schober, A. ,Leng, L.
(2007) MIF is a noncognate ligand of CXC chemokine recep-
tors in in ammatory and atherogenic cell recruitment. Nature
medicine, Vol. 13, No. 5, pp. 587-96.
Bozek, K., Lengauer, T., Sierra, S., Kaiser, R. ,Domingues, F.S.
(2013) Analysis of physicochemical and structural properties
determining HIV-1 coreceptor usage. PLOS Comput Biol, Vol.
9, No. 3, p. e1002977.
Checkley, M.A., Luttge, B.G. ,Freed, E.O. (2011) HIV-1 enve-
lope glycoprotein biosynthesis, traf cking, and incorporation.
Journal of molecular biology, Vol. 410, No. 4, pp. 582-608.
Choi, I., Chung, A.W., Suscovich, T.J., Rerks-Ngarm, S., Pit-
isuttithum, P., Nitayaphan, S., Kaewkungwal, J., O’Connell,
R.J., Francis, D. ,Robb, M.L. (2015) Machine learning methods
enable predictive modeling of antibody feature: function rela-
tionships in RV144 vaccinees. PLOS Comput Biol, Vol. 11, No.
4, p. e1004185.
Cutlan, J., Saunders, N., Olsen, S. ,Fullen, D. (2010) White sponge
nevus presenting as genital lesions in a 28‐year‐old female.
Journal of cutaneous pathology, Vol. 37, No. 3, pp. 386-9.
Daniels, V.G. (2012) AIDS: the acquired immune de ciency
syndrome, Springer Science & Business Media.
Dawany, N., Showe, L.C., Kossenkov, A.V., Chang, C., Ive, P.,
Conradie, F., Stevens, W., Sanne, I., Azzoni, L. ,Montaner, L.J.
(2014) Identi cation of a 251 gene expression signature that
can accurately detect M. tuberculosis in patients with and
without HIV co-infection. PloS one, Vol. 9, No. 2, p. e89925.
De Silva, E. ,Stumpf, M.P. (2004) HIV and the CCR5-Δ32 resist-
ance allele. FEMS microbiology letters, Vol. 241, No. 1, pp. 1-12.
Di Benedetto, F., Di Sandro, S., De Ruvo, N., Berretta, M.,
Masetti, M., Montalti, R., Ballarin, R., Cocchi, S., Potenza, L.
,Luppi, M. (2008) Kaposi’s sarcoma after liver transplantation.
Journal of cancer research and clinical oncology, Vol. 134, No.
6, pp. 653-8.
Doitsh, G., Cavrois, M., Lassen, K.G., Zepeda, O., Yang, Z., San-
tiago, M.L., Hebbeler, A.M. ,Greene, W.C. (2010) Abortive HIV
infection mediates CD4 T cell depletion and in ammation in
human lymphoid tissue. Cell, Vol. 143, No. 5, pp. 789-801.
Dybowski, J.N., Heider, D. ,Hoffmann, D. (2010) Prediction of
co-receptor usage of HIV-1 from genotype. PLOS Comput Biol,
Vol. 6, No. 4, p. e1000743.
BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS MACHINE LEARNING APPROACHES TO STUDY HIV/AIDS INFECTION 41

Sweta, Usha and Sunil
Evans, M.C., Paquet, A.C., Huang, W., Napolitano, L., Frant-
zell, A., Toma, J., Stawiski, E.W., Goetz, M.B., Petropoulos,
C.J. ,Whitcomb, J. (2013) A case-based reasoning system for
genotypic prediction of hiv-1 co-receptor tropism. Journal of
bioinformatics and computational biology, Vol. 11, No. 04, p.
1350006.
Evans, M.C., Phung, P., Paquet, A.C., Parikh, A., Petropoulos,
C.J., Wrin, T. ,Haddad, M. (2014) Predicting HIV-1 broadly
neutralizing antibody epitope networks using neutralization
titers and a novel computational method. Bmc Bioinformatics,
Vol. 15, No. 1, p. 77.
Fogel, G.B., Lamers, S.L., Liu, E.S., Salemi, M. ,McGrath, M.S.
(2015) Identi cation of dual-tropic HIV-1 using evolved neural
networks. Biosystems, Vol. 137, pp. 12-9.
Gorry, P.R., Churchill, M., Crowe, S.M., Cunningham, A.L.
,Gabuzda, D. (2005) Pathogenesis of macrophage tropic HIV-1.
Current HIV research, Vol. 3, No. 1, pp. 53-60.
Gupta, S., Neogi, U., Srinivasa, H. ,Shet, A. (2015) Performance
of genotypic tools for prediction of tropism in HIV-1 subtype C
V3 loop sequences. Intervirology, Vol. 58, No. 1, pp. 1-5.
Heider, D., Senge, R., Cheng, W. ,Hüllermeier, E. (2013) Multi-
label classi cation for exploiting cross-resistance information
in HIV-1 drug resistance prediction. Bioinformatics, Vol. 29,
No. 16, pp. 1946-52.
Hepler, N.L., Schef er, K., Weaver, S., Murrell, B., Richman,
D.D., Burton, D.R., Poignard, P., Smith, D.M. ,Pond, S.L.K.
(2014) IDEPI: rapid prediction of HIV-1 antibody epitopes and
other phenotypic features from sequence data using a exible
machine learning platform. PLOS Comput Biol, Vol. 10, No. 9,
p. e1003842.
Holman, A.G. ,Gabuzda, D. (2012) A machine learning approach
for identifying amino acid signatures in the HIV env gene pre-
dictive of dementia. PloS one, Vol. 7, No. 11, p. e49538.
Hütter, G., Nowak, D., Mossner, M., Ganepola, S., Müßig, A.,
Allers, K., Schneider, T., Hofmann, J., Kücherer, C. ,Blau, O.
(2009) Long-term control of HIV by CCR5 Delta32/Delta32
stem-cell transplantation. New England Journal of Medicine,
Vol. 360, No. 7, pp. 692-8.
Ji, C., Zhang, J., Dioszegi, M., Chiu, S., Rao, E., Cammack, N.,
Brandt, M. ,Sankuratri, S. (2007) CCR5 small-molecule antago-
nists and monoclonal antibodies exert potent synergistic anti-
viral effects by cobinding to the receptor. Molecular pharma-
cology, Vol. 72, No. 1, pp. 18-28.
Kaplan, J.E., Benson, C., Holmes, K.K., Brooks, J.T., Pau, A.,
Masur, H., Control, C.f.D., Prevention, Health, N.I.o. ,America,
H.M.A.o.t.I.D.S.o. (2009) Guidelines for prevention and treat-
ment of opportunistic infections in HIV-infected adults and
adolescents. MMWR Recomm Rep, Vol. 58, No. RR-4, pp.
1-207.
Kay, M.A., Walker, B.D. (2014) Engineering cellular resistance
to HIV. N Engl J Med, Vol. 370, No. 10, pp. 968-9.
Kim, G., Kim, Y., Lim, H. ,Kim, H. (2010) An MLP-based feature
subset selection for HIV-1 protease cleavage site analysis. Arti-
cial intelligence in medicine, Vol. 48, No. 2, pp. 83-9.
Kirchmair, J., Distinto, S., Roman Liedl, K., Markt, P., Maria
Rollinger, J., Schuster, D., Maria Spitzer, G. ,Wolber, G. (2011)
Development of anti-viral agents using molecular modeling
and virtual screening techniques. Infectious Disorders-Drug
Targets (Formerly Current Drug Targets-Infectious Disorders),
Vol. 11, No. 1, pp. 64-93.
Kumar, R., Raghava, G.P. (2013) Hybrid approach for predict-
ing coreceptor used by HIV-1 from its V3 loop amino acid
sequence. PloS one, Vol. 8, No. 4, p. e61437.
Majetschak, M. (2011) Extracellular ubiquitin: immune modu-
lator and endogenous opponent of damage-associated molecu-
lar pattern molecules. Journal of leukocyte biology, Vol. 89,
No. 2, pp. 205-19.
Masso, M., Vaisman, I.I. (2010) Accurate and ef cient gp120
V3 loop structure based models for the determination of HIV-1
co-receptor usage. Bmc Bioinformatics, Vol. 11, No. 1, p. 494.
Mathevula, H.M. (2013) Factors affecting adherence to treat-
ment in patients on chronic medication at Mokopane Hospital,
University of Limpopo (Tur oop Campus).
Mathewos, M. (2014) Prevalence of opportunistic intestinal
parasitic infections among hiv/aids patients attending Othona
hospital, Wolayita Sodo, Southern Ethiopia.
Mei, S. (2013) Probability weighted ensemble transfer learning
for predicting interactions between HIV-1 and human proteins.
PloS one, Vol. 8, No. 11, p. e79606.
Miyakawa, T., Obaru, K., Maeda, K., Harada, S. ,Mitsuya, H.
(2002) Identi cation of amino acid residues critical for LD78,
a variant of human macrophage in ammatory protein-1,
binding to CCR5 and inhibition of R5 human immunode -
ciency virus type 1 replication. Journal of Biological Chemis-
try, Vol. 277, No. 7, pp. 4649-55.
Murphy, P.M. (2001) Viral exploitation and subversion of the
immune system through chemokine mimicry. Nature immu-
nology, Vol. 2, No. 2, pp. 116-22.
Nakayama, K., Nakamura, H., Koga, M., Koibuchi, T., Fujii, T.,
Miura, T., Iwamoto, A. ,Kawana-Tachikawa, A. (2012) Imbal-
anced production of cytokines by T cells associates with the
activation/exhaustion status of memory T cells in chronic HIV
type 1 infection. AIDS research and human retroviruses, Vol.
28, No. 7, pp. 702-14.
Öztürk, O., Aksaç, A., Elsheikh, A., Özyer, T. ,Alhajj, R. (2013) A
consistency-based feature selection method allied with linear
SVMs for HIV-1 protease cleavage site prediction. PloS one,
Vol. 8, No. 8, p. e63145.
Page, J., Louw, M. ,Pakkiri, D. 2006, Working with HIV/Aids,
Juta and Company Ltd.
Pancholi, N.J., Gupta, S., Sapre, N. ,Sapre, N.S. (2014) Design
of novel leads: ligand based computational modeling stud-
ies on non-nucleoside reverse transcriptase inhibitors
(NNRTIs) of HIV-1. Molecular BioSystems, Vol. 10, No. 2,
pp. 313-25.
Park, B.J., Wannemuehler, K.A., Marston, B.J., Govender, N.,
Pappas, P.G. ,Chiller, T.M. (2009) Estimation of the current
42 MACHINE LEARNING APPROACHES TO STUDY HIV/AIDS INFECTION BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS

Sweta, Usha and Sunil
global burden of cryptococcal meningitis among persons liv-
ing with HIV/AIDS. Aids, Vol. 23, No. 4, pp. 525-30.
Pavlasova, G., Borsky, M., Seda, V., Cerna, K., Osickova, J.,
Doubek, M., Mayer, J., Calogero, R., Trbusek, M. ,Pospisilova,
S. (2016) Ibrutinib inhibits CD20 upregulation on CLL B cells
mediated by the CXCR4/SDF-1 axis. Blood, Vol. 128, No. 12,
pp. 1609-13.
Pawlowski, A., Jansson, M., Sköld, M., Rottenberg, M.E. ,Käl-
lenius, G. (2012) Tuberculosis and HIV co-infection. PLoS Pat-
hog, Vol. 8, No. 2, p. e1002464.
Poongavanam, V., Kongsted, J. (2013) Virtual screening models
for prediction of HIV-1 RT associated RNase H inhibition. PloS
one, Vol. 8, No. 9, p. e73478.
Prosperi, M., De Luca, A. (2011) Computational models for pre-
diction of response to antiretroviral therapies. AIDS reviews,
Vol. 14, No. 2, pp. 145-53.
Prosperi, M.C., Altmann, A., Rosen-Zvi, M., Aharoni, E., Bor-
gulya, G., Bazso, F., Sönnerborg, A., Schülter, E., Struck, D.
,Ulivi, G. (2009) Investigation of expert rule bases, logistic
regression, and non-linear machine learning techniques for
predicting response to antiretroviral treatment. Antivir Ther,
Vol. 14, No. 3, pp. 433-42.
Riemenschneider, M., Heider, D. (2016a) Current Approaches in
Computational Drug Resistance Prediction in HIV. Current HIV
research, Vol. 14, No. 4, pp. 307-15.
Riemenschneider, M., Senge, R., Neumann, U., Hüllermeier,
E. ,Heider, D. (2016b) Exploiting HIV-1 protease and reverse
transcriptase cross-resistance information for improved drug
resistance prediction by means of multi-label classi cation.
BioData mining, Vol. 9, No. 1, p. 10.
Saini, V., Marchese, A. ,Majetschak, M. (2010) CXC chemokine
receptor 4 is a cell surface receptor for extracellular ubiquitin.
Journal of Biological Chemistry, Vol. 285, No. 20, pp. 15566-
76.
Samson, M., Labbe, O., Mollereau, C., Vassart, G. ,Parmentier,
M. (1996) Molecular cloning and functional expression of a
new human CC-chemokine receptor gene. Biochemistry, Vol.
35, No. 11, pp. 3362-7.
Sander, O., Sing, T., Sommer, I., Low, A.J., Cheung, P.K., Har-
rigan, P.R., Lengauer, T. ,Domingues, F.S. (2007) Structural
descriptors of gp120 V3 loop for the prediction of HIV-1 core-
ceptor usage. PLOS Comput Biol, Vol. 3, No. 3, p. e58.
Scherzer, R., Estrella, M., Yongmei, L., Deeks, S.G., Grunfeld, C.
,Shlipak, M.G. (2012) Association of tenofovir exposure with
kidney disease risk in HIV infection. AIDS (London, England),
Vol. 26, No. 7, p. 867.
Singh, O., Su, E.C.-Y. (2016) Prediction of HIV-1 protease
cleavage site using a combination of sequence, structural, and
physicochemical features. Bmc Bioinformatics, Vol. 17, No. 17,
p. 279.
Slimani, H., Charnaux, N., Mbemba, E., Saffar, L., Vassy, R.,
Vita, C. ,Gattegno, L. (2003) Interaction of RANTES with syn-
decan-1 and syndecan-4 expressed by human primary mac-
rophages. Biochimica et Biophysica Acta (BBA)-Biomem-
branes, Vol. 1617, No. 1, pp. 80-8.
Srisawat, A., Kijsirikul, B. (2008) Combining classi ers for
HIV-1 drug resistance prediction. Protein and peptide letters,
Vol. 15, No. 5, pp. 435-42.
Struyf, S., Menten, P., Lenaerts, J.P., Put, W., D’Haese, A., De
Clercq, E., Schols, D., Proost, P. ,Van Damme, J. (2001) Diverg-
ing binding capacities of natural LD78 isoforms of mac-
rophage in ammatory protein‐1 to the CC chemokine recep-
tors 1, 3 and 5 affect their anti‐HIV‐1 activity and chemotactic
potencies for neutrophils and eosinophils. European journal of
immunology, Vol. 31, No. 7, pp. 2170-8.
Tebas, P., Stein, D., Tang, W.W., Frank, I., Wang, S.Q., Lee, G.,
Spratt, S.K., Surosky, R.T., Giedlin, M.A. ,Nichol, G. (2014)
Gene editing of CCR5 in autologous CD4 T cells of persons
infected with HIV. New England Journal of Medicine, Vol. 370,
No. 10, pp. 901-10.
Wawer, M.J., Gray, R.H., Sewankambo, N.K., Serwadda, D., Li,
X., Laeyendecker, O., Kiwanuka, N., Kigozi, G., Kiddugavu, M.
,Lutalo, T. (2005) Rates of HIV-1 transmission per coital act, by
stage of HIV-1 infection, in Rakai, Uganda. Journal of Infec-
tious Diseases, Vol. 191, No. 9,
pp. 1403-9.
Wei, Y., Li, J., Chen, Z., Wang, F., Huang, W., Hong, Z. ,Lin, J.
(2015) Multistage virtual screening and identi cation of novel
HIV-1 protease inhibitors by integrating SVM, shape, pharma-
cophore and docking methods. European journal of medicinal
chemistry, Vol. 101, pp. 409-18.
Wiskerchen, M. ,Muesing, M.A. (1995) Human immunode -
ciency virus type 1 integrase: effects of mutations on viral
ability to integrate, direct viral gene expression from unin-
tegrated viral DNA templates, and sustain viral propaga-
tion in primary cells. Journal of virology, Vol. 69, No. 1,
pp. 376-86.
Xuan, S., Wu, Y., Chen, X., Liu, J. ,Yan, A. (2013) Prediction
of bioactivity of HIV-1 integrase ST inhibitors by multilinear
regression analysis and support vector machine. Bioorganic &
medicinal chemistry letters, Vol. 23, No. 6, pp. 1648-55.
Yan, A., Xuan, S. ,Hu, X. (2012) Classi cation of active and
weakly active ST inhibitors of HIV-1 integrase using a support
vector machine. Combinatorial chemistry & high throughput
screening, Vol. 15, No. 10, pp. 792-805.
Zazzi, M., Cozzi-Lepri, A. ,Prosperi, M.C. (2016) Computer-
Aided Optimization of Combined Anti-Retroviral Therapy for
HIV: New Drugs, New Drug Targets and Drug Resistance. Cur-
rent HIV research, Vol. 14, No. 2, pp. 101-9.
Zazzi, M., Incardona, F., Rosen-Zvi, M., Prosperi, M., Lengauer,
T., Altmann, A., Sonnerborg, A., Lavee, T., Schülter, E. ,Kai-
ser, R. (2012) Predicting response to antiretroviral treatment by
machine learning: the EuResist project. Intervirology, Vol. 55,
No. 2, pp. 123-7.
Zhen, A. ,Kitchen, S. (2013) Stem-cell-based gene therapy for
HIV infection. Viruses, Vol. 6, No. 1, pp. 1-12.
BIOSCIENCE BIOTECHNOLOGY RESEARCH COMMUNICATIONS MACHINE LEARNING APPROACHES TO STUDY HIV/AIDS INFECTION 43